The inla.posterior.sample.eval-function
Haavard Rue
KAUST, Aug 2022
sample-eval.RmdIntroduction
This short note add some more explanation to the
inla.posterior.sample.eval()-function, as its a constant
source of confusing (which is understandable). The purpose of this
function is to ease function-evaluations of samples from the fitted
model.
Simple example
As often, its easier to work with an example.
n <- 100
x <- rnorm(n)
eta <- 1 + x
y <- rnorm(n, mean = eta, sd = 0.1)
r <- inla(y ~ 1 + x,
data = data.frame(y,x),
control.compute = list(config=TRUE))where the config argument is required. We can now generate samples from the fitted model
samples <- inla.posterior.sample(10000, r)Here, samples contains the samples, but in long vectors with additional information about where to find what which makes it complicated.
The information available is what is in the output
summary(r)Time used:
Pre = 0.354, Running = 0.157, Post = 0.0353, Total = 0.546
Fixed effects:
mean sd 0.025quant 0.5quant 0.975quant mode kld
(Intercept) 0.990 0.010 0.971 0.990 1.009 0.990 0
x 0.995 0.011 0.974 0.995 1.016 0.995 0
Model hyperparameters:
mean sd 0.025quant 0.5quant
Precision for the Gaussian observations 108.27 15.31 80.39 107.55
0.975quant mode
Precision for the Gaussian observations 140.30 106.11
Marginal log-Likelihood: 74.36
is computed
Posterior summaries for the linear predictor and the fitted values are computed
(Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')so its x, (Intercept) and the Precision for the Gaussian observations, plus the linear predictor(s).
The ...eval() function simplifies the evaluation of a
function over (joint) samples, by assigning sample-values to each
variable. To extract samples of x, which is here the
regression coefficient for the covariates
(this is confusing, I know), then we can do
fun1 <- function() return (x)We can now evaluate fun1 for each sample, using the
...eval()-function, like
eval.fun1 <- inla.posterior.sample.eval(fun1, samples)
str(eval.fun1) num [1, 1:10000] 1.006 0.97 0.977 0.998 1.004 ...
- attr(*, "dimnames")=List of 2
..$ : chr "fun[1]"
..$ : chr [1:10000] "sample:1" "sample:2" "sample:3" "sample:4" ...since x is automatically assigned the sample value
before fun1 is called. This happens for each sample.
We can compare with the the INLA-output
hist(eval.fun1[1,], prob=TRUE, n=300)
lines(inla.smarginal(r$marginals.fixed$x), lwd=3)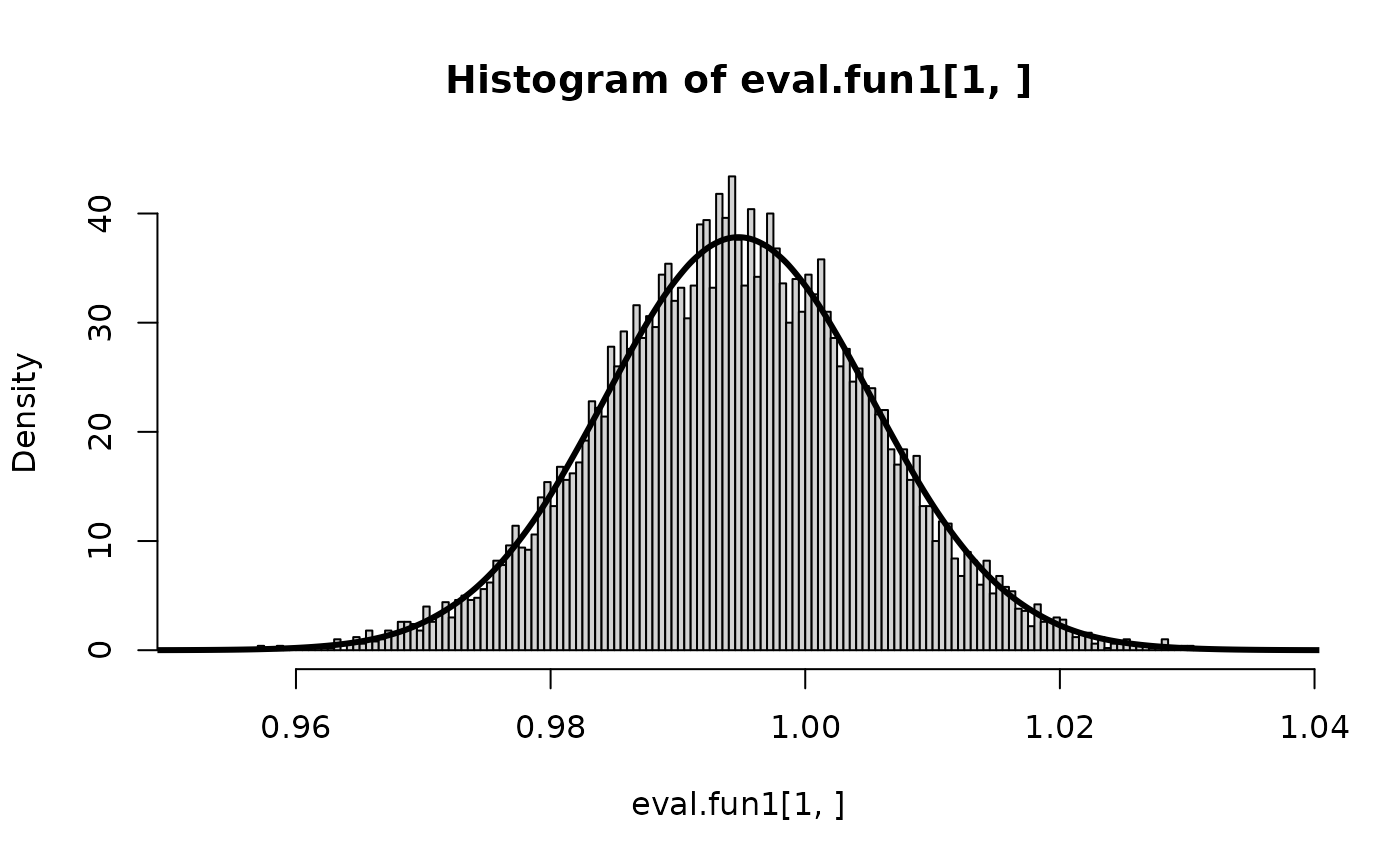 and the results seems to agree.
and the results seems to agree.
Also the variable (Intercept) is automatically created, but since this form is awkward to use in R, it is equivalent to Intercept. We can for example sample the linear predictor with
fun2 <- function(x.cov) return (Intercept + x * x.cov)Here, we need to pass the covariates (which is not the same as x) separately as a named argument,
eval.fun2 <- inla.posterior.sample.eval(fun2, samples, x.cov = x)and we plot the regression-line
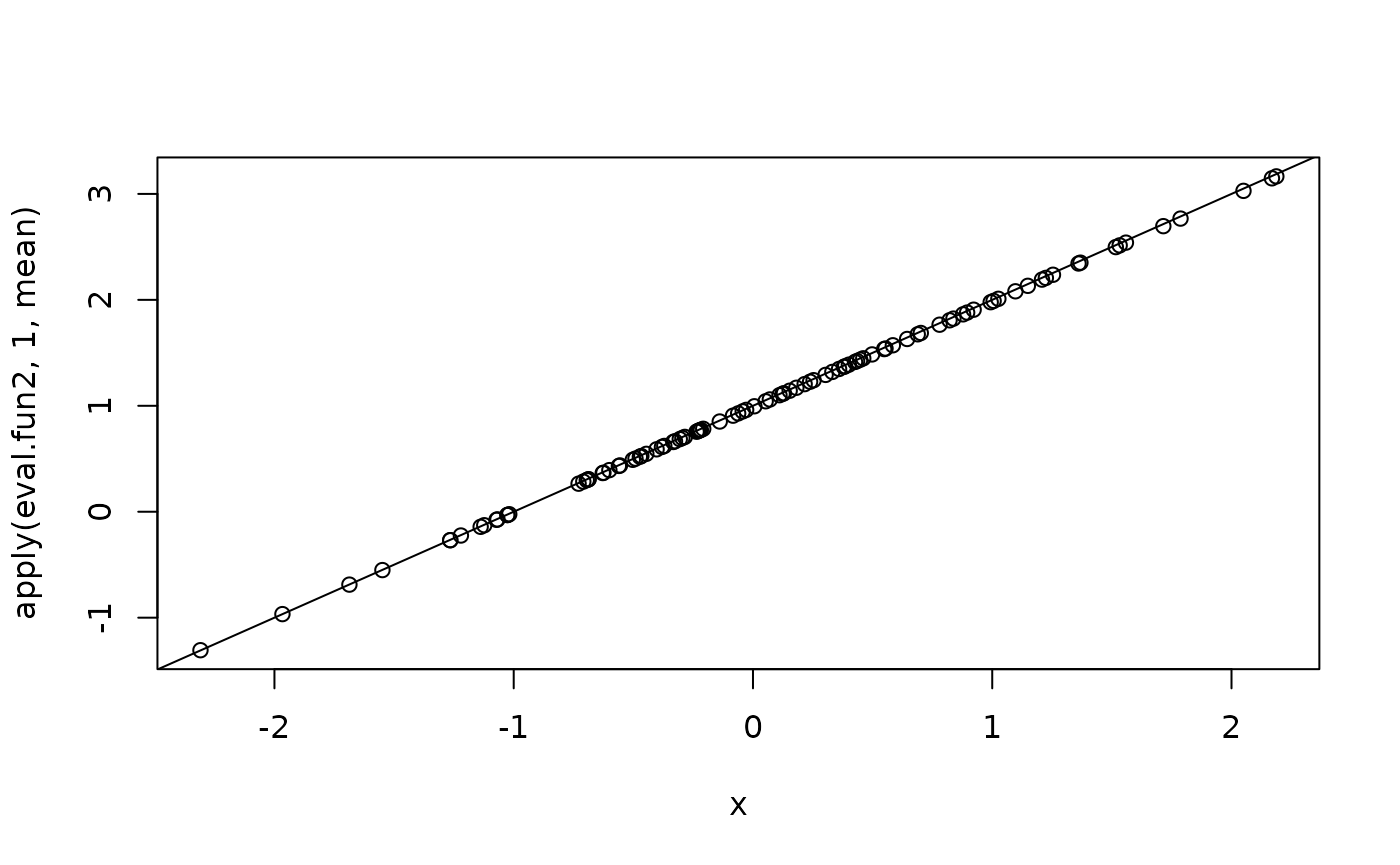 The predictor is also available automatically as
Predictor, so
The predictor is also available automatically as
Predictor, so
fun3 <- function(x.cov) return (Predictor - (Intercept + x * x.cov))
eval.fun3 <- inla.posterior.sample.eval(fun3, samples, x.cov = x)
summary(eval.fun3[1,]) Min. 1st Qu. Median Mean 3rd Qu. Max.
-5.551e-17 -5.551e-17 0.000e+00 -4.940e-19 0.000e+00 5.551e-17 as it should.
Samples of hyper-parameters
It gets a little more involved with the hyper-parameters. In the example above, there is only one, the precision for the observational noise. We can use this to sample new data from the fitted model. Hyper-parameters are automatically assigned values in the vector theta.
fun4 <- function() return (theta)
eval.fun4 <- inla.posterior.sample.eval(fun4, samples)
table(eval.fun4[1, ])
61.6806477523926 72.4386693479479 81.7215542045142 92.1940240166842
11 93 541 1452
99.911053924203 108.274031887778 116.73423974298 125.855502845737
1969 1915 2031 1435
140.890961882934 157.722647730621
505 48 A feature here, is that only the integration points for
theta are used, hence samples of theta
are discrete with finite number of values. (To only sample the
hyper-parameters, please use function
inla.hyperpar.sample().) Note that theta,
by default, is the hyper-parameters in the user-scale (like precision,
correlation, etc). If argument intern=TRUE is used in the
inla.posterior.sample()-function, then they will appear in
the internal-scale (like log(precision), etc).
We can generate a new dataset from the fitted model, with
samples <- inla.posterior.sample(1, r)
fun4 <- function() {
n <- length(Predictor)
return (Predictor + rnorm(n, sd = sqrt(1/theta)))
}
eval.fun4 <- inla.posterior.sample.eval(fun4, samples)
plot(x, eval.fun4[,1])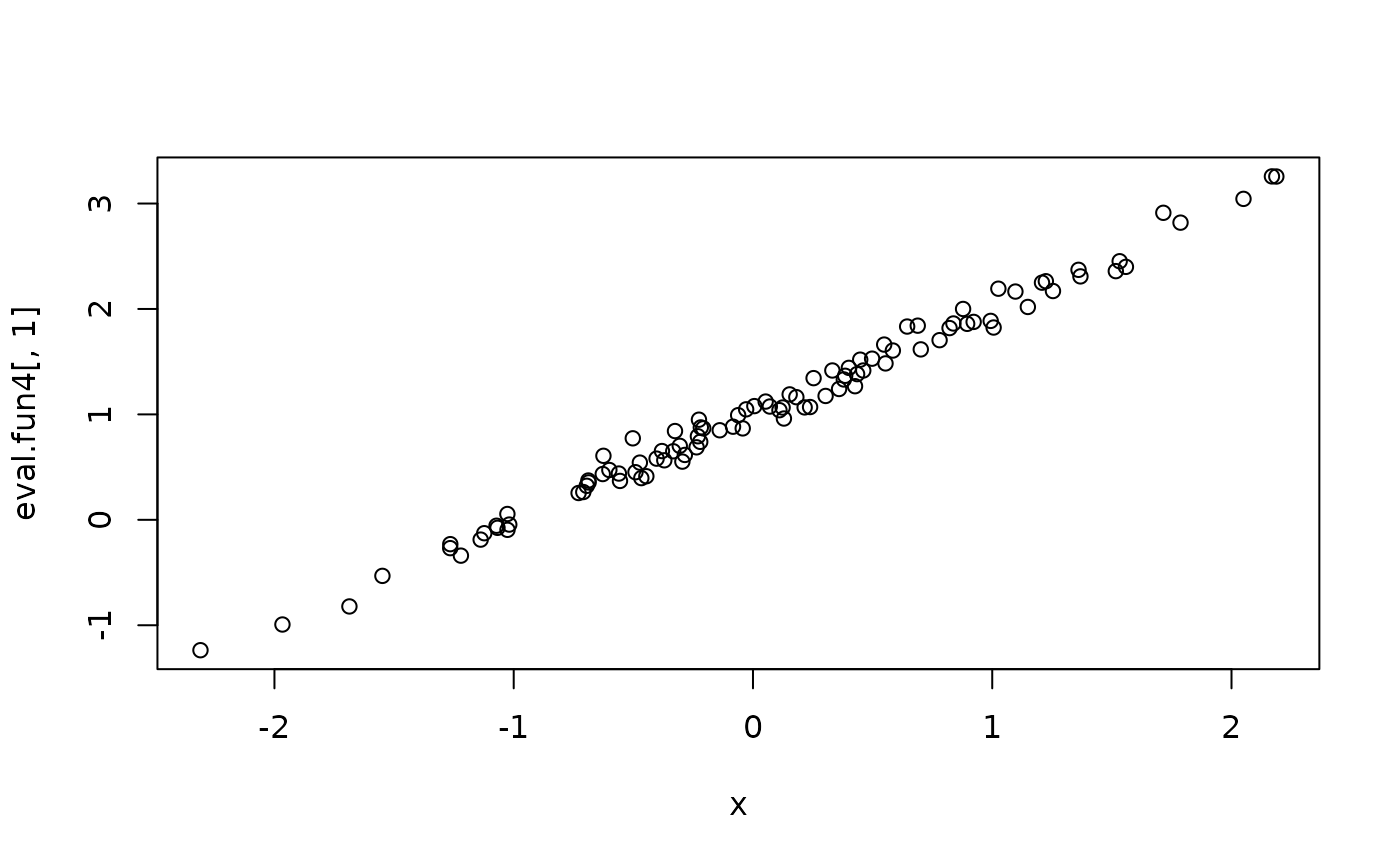
With more than one hyper-parameter, then theta is
vector, and the order of hyper-parameters is the same as is stored in
the result-object. The user has to organise this manually. With
r <- inla(y ~ 1 + x,
family = "sn",
data = data.frame(y,x),
control.compute = list(config=TRUE))then
rownames(r$summary.hyperpar)[1] "precision for skew-normal observations"
[2] "Skewness for skew-normal observations" so that theta[1] is the precision while
theta[2] is the skewness.
Example: Predictor with and without random effects
Here is an example that pops up from time to time, using the tools above. We are interested in comparing the linear predictor with and without some random effects. The below example is artificial but shows how this works.
First we simulate some data
m <- 100
n <- m^2
## fixed effects
x <- rnorm(n)
xx <- rnorm(n)
## random effects
v <- rnorm(m, sd=0.2)
v.idx <- rep(1:m, each = m)
eta <- 1 + 0.2 * (x + xx) + v
y <- rpois(n, exp(eta))and then fit the model
r <- inla(y ~ 1 + x + xx + f(v.idx, model = "iid"),
data = data.frame(y, x, xx, v.idx),
family = "poisson",
control.compute = list(config = TRUE))
samples <- inla.posterior.sample(10000, r)Now we want to compare the linear predictor with and without the
f(v.idx, model = "iid") term. The easy way out, is to use
the predictor and then subtract the iid-term, instead of building it up
manually.
fun5 <- function(v.index) {
return (c(Predictor, Predictor - v.idx[v.index]))
}
eval.fun5 <- inla.posterior.sample.eval(fun5, samples, v.index=v.idx)And we can compare with and without for some components
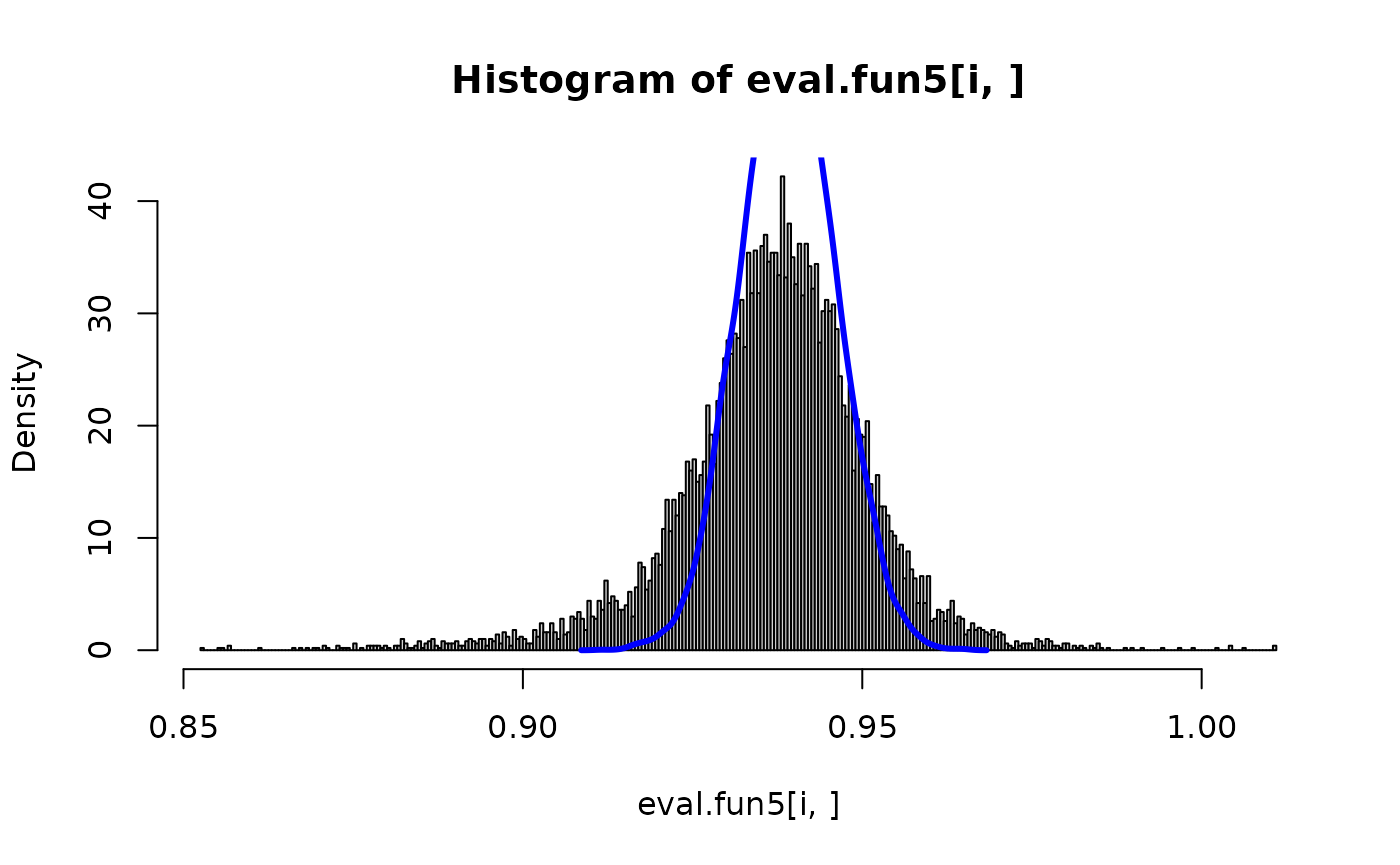
Predictor -matrix (experimental-mode only)
For some models, especially models using the SPDE, then a projector matrix is used, so we need the A-matrix for the predictor. Often this looks like
r <- inla(...., control.predictor = list(A=inla.stack.A(...)))For these models, then the observations depend on
,
where
,
and
is defined with the formula. In these cases, then Predictor
is
and APredictor is
.
Moreover, the
matrix is available as pA in the
...eval()-function.
Here is the same example above, with a random -matrix showing how to use this.
n <- 100
m <- 25
## fixed effects
x <- rnorm(n)
xx <- rnorm(n)
## random effects
v <- rnorm(m, sd=0.2)
v.idx <- rep(1:m, each = n %/% m)
eta <- 1 + 0.2 * (x + xx) + v
A <- matrix(rnorm(n^2, sd=sqrt(1/n)), n, n)
eta.star <- A %*% eta
y <- rpois(n, exp(eta.star))
r <- inla(y ~ 1 + x + xx + f(v.idx, model = "iid"),
data = data.frame(y, x, xx, v.idx),
family = "poisson",
inla.mode = "experimental",
control.predictor = list(A=A),
control.compute = list(config = TRUE))
samples <- inla.posterior.sample(10000, r)We will compare the same change, with and without the iid-term.
fun6 <- function(v.index) {
return (c(APredictor, as.numeric(pA %*% (Predictor - v.idx[v.index]))))
}
eval.fun6 <- inla.posterior.sample.eval(fun6, samples, v.index=v.idx)
i <- 2
hist(eval.fun6[i,], prob=TRUE, n=300)
lines(density(eval.fun6[n + i,]), col="blue", lwd=3)