A note about values
Håvard Rue (hrue@r-inla.org) & Sigrunn Holbek Sørbye (sigrunn.sorbye@uit.no)
Dec, 2025
a-note-about-values.RmdIntroduction
This a short note discussing the argument in , to avoid unfortunate confusion. This is especially an issue for models like , and , which are only defined for integer-valued arguments.
Consider the AR1-model for .
In practice, we might use the AR1-model for various sequential indexes, like years, weeks, etc. In these cases, it is naturally to write
y ~ ... + f(year, model = "ar1", ...) where for example is given by
year <- 2000:2025What happens internally, is that the -variable specifies the (time points) at which the AR1-model is defined. So, the variable would be mapped to the -index, which runs from to .
To be more specific, a mapping between and the -index is defined as
## [1] 2000 2001 2002 2003 2004 2005 2006 2007 2008 2009 2010 2011 2012 2013 2014
## [16] 2015 2016 2017 2018 2019 2020 2021 2022 2023 2024 2025
map.year.to.t <- function(year, values) {
a <- numeric(length(year))
for(i in seq_along(year))
a[i] = which(year[i] == values)
return (a)
}
yy <- c(2000, 2020, 2025)
print(cbind(year=yy, t=map.year.to.t(yy, values)))## year t
## [1,] 2000 1
## [2,] 2020 21
## [3,] 2025 26An unfortunate consequence is that if some years are missing, then “confusing” things happen. Suppose, we do not have obsevations for some years,
year <- c(2000:2020, 2023:2025)Then the above mapping still applies but with
## [1] 2000 2001 2002 2003 2004 2005 2006 2007 2008 2009 2010 2011 2012 2013 2014
## [16] 2015 2016 2017 2018 2019 2020 2023 2024 2025in which the years 2021 and 2022 are missing. The internal mapping now gives
## year t
## [1,] 2000 1
## [2,] 2020 21
## [3,] 2025 24Note that for the index is now 24 and not 26 as earlier. This implies that the correlation between year 2020 and 2023 is while it was as before. Often, this is not what we expect to happen.
Another case is when the argument denote non-integers, like
time <- c(1.1, 2.2, 3.3, 4.4, 10.0)A call using would run, but will be mapped to the discrete indexes . This is because the models are defined on integers and cannot handle irregular locations.
We can ensure consistent behaviour by adding the argument to , specifying for which indexes the model is defined, like
y ~ ... + f(year, model = "ar1", values = 2000:2025)or, if we want predictions for additional years,
y ~ ... + f(year, model = "ar1", values = 1990:2035)In this case, the are defined by the corresponding named argument and not by the variable as before
The requirement for is now that these are either -values or a subset of the defined .
An example
This is a simple simulated example demonstrating the issue about , having missing observations.
year <- 2000:2025
n <- length(year)
phi <- 0.9
phi.intern <- inla.models()$latent$ar1$hyper$theta2$to.theta(phi)
x <- scale(arima.sim(n, model=list(ar=phi)))
s <- 0.3
y <- x + rnorm(n, sd=s)
plot(year, y, pch=19)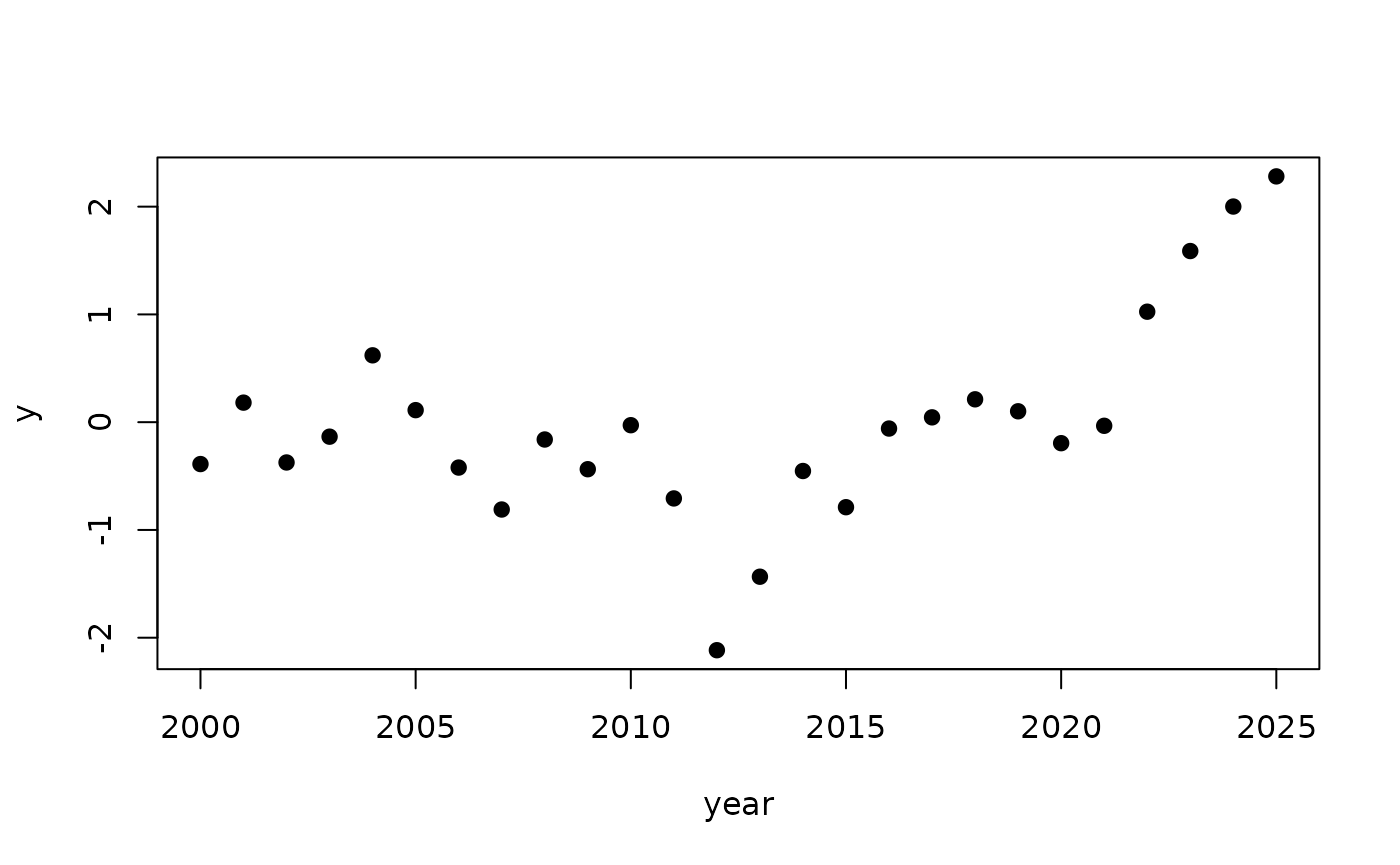
Let us fit an AR1 model
r <- inla(y ~ -1 + f(year, model="ar1",
hyper = list(prec = list(initial=0,
fixed=TRUE),
rho = list(initial=phi.intern,
fixed=TRUE))),
family="gaussian",
control.family = list(hyper = list(
prec = list(initial=log(1/s^2),
fixed = TRUE))),
data = data.frame(y, year))
plot(year, y, pch=19)
lines(year, r$summary.random$year$mean)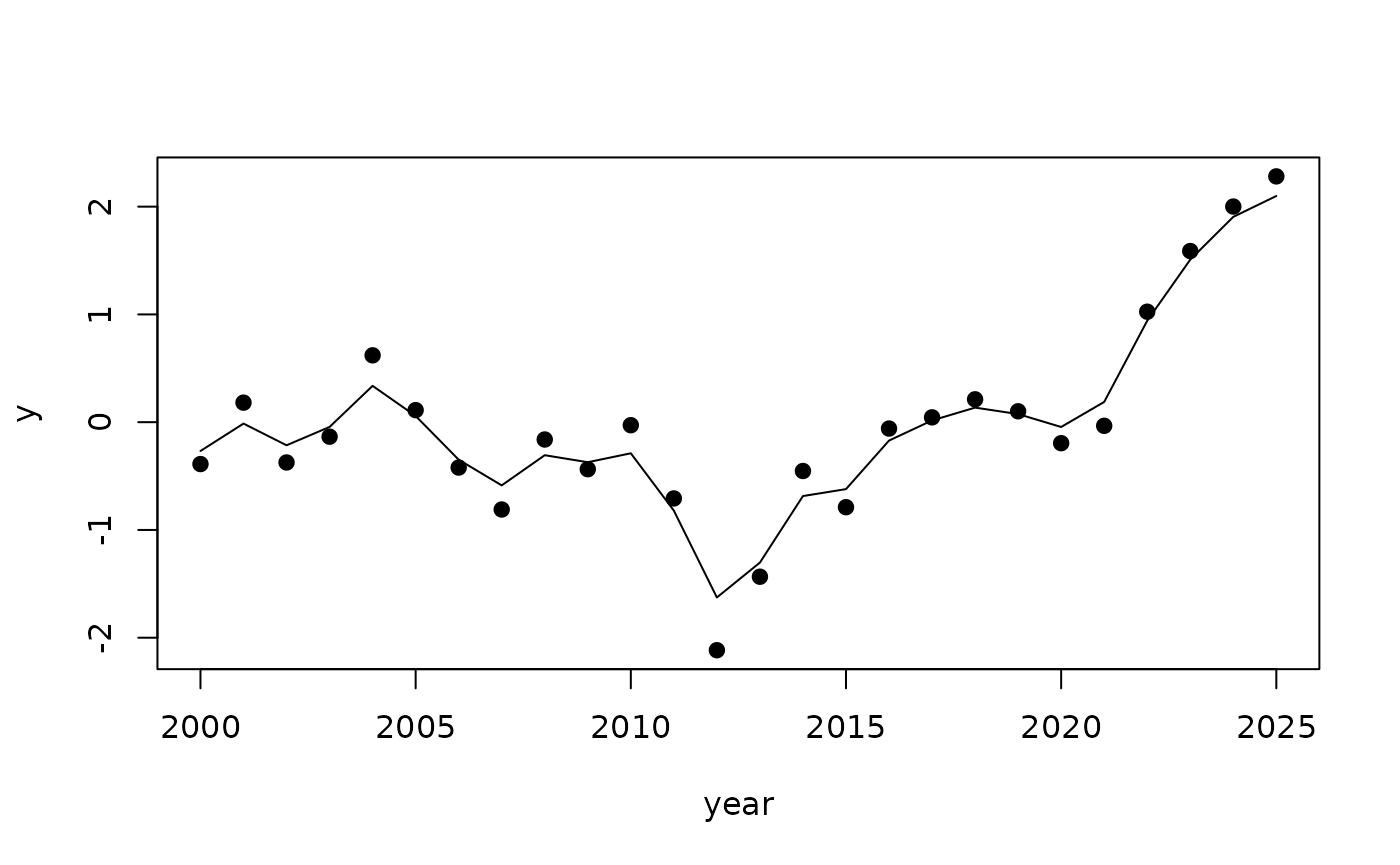
Assume is missing, then we will get the wrong results by removing from the data, because are not given (hence is removed as well).
## this is correct
y[10] <- NA
r <- inla(y ~ -1 + f(year, model="ar1",
hyper = list(prec = list(initial=0,
fixed=TRUE),
rho = list(initial=phi.intern,
fixed=TRUE))),
family="gaussian",
control.family = list(hyper = list(
prec = list(initial=log(1/s^2),
fixed = TRUE))),
data = data.frame(y, year)) ## missing data included
## this is wrong
rr <- inla(y ~ -1 + f(year, model="ar1",
hyper = list(prec = list(initial=0,
fixed=TRUE),
rho = list(initial=phi.intern,
fixed=TRUE))),
family="gaussian",
control.family = list(hyper = list(
prec = list(initial=log(1/s^2),
fixed = TRUE))),
data = data.frame(y, year)[-10,]) ## missing data removed
## we compare the results
r$mlik - rr$mlik## [,1]
## log marginal-likelihood (integration) -0.1904075
## log marginal-likelihood (Gaussian) -0.1904075If we add , then we get the correct result when we remove missing data.
rrr <- inla(y ~ -1 + f(year, model="ar1",
values = 2000:2025, ## values added
hyper = list(prec = list(initial=0,
fixed=TRUE),
rho = list(initial=phi.intern,
fixed=TRUE))),
family="gaussian",
control.family = list(hyper = list(
prec = list(initial=log(1/s^2),
fixed = TRUE))),
data = data.frame(y, year)[-10,]) ## missing data removed
## we compare the results
r$mlik - rrr$mlik## [,1]
## log marginal-likelihood (integration) 2.842171e-14
## log marginal-likelihood (Gaussian) 2.842171e-14Another example
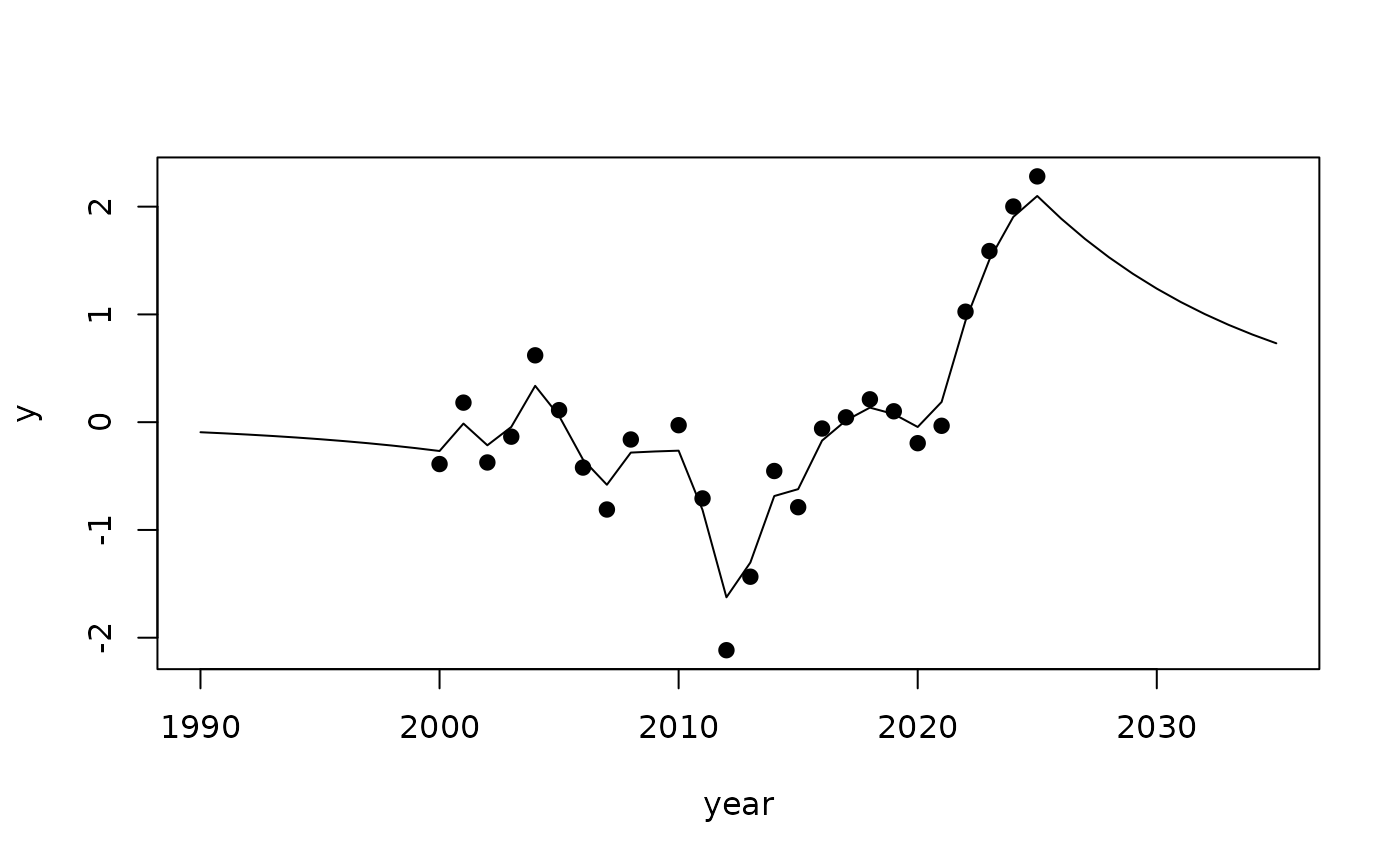
We can extend the range using , to get predictions.
values <- 1990:2035
r <- inla(y ~ -1 + f(year, model="ar1",
values = values, ## values added
hyper = list(prec = list(initial=0,
fixed=TRUE),
rho = list(initial=phi.intern,
fixed=TRUE))),
family="gaussian",
control.family = list(hyper = list(
prec = list(initial=log(1/s^2),
fixed = TRUE))),
data = list(y=y, year=year, values=values))
plot(year, y, pch=19, xlim=range(values))
lines(values, r$summary.random$year$mean)