User defined integration points
Haavard Rue
(hrue@r-inla.org)
May 2017 (updated April 2019 and May 2023)
int-design.RmdIntroduction
This short note describe a new option that allow the user to use user-defined integration points (or “design” points), instead of the default ones. The relevant integration in INLA does where is the approximated posterior marginal for the hyperparameters, and where is the approximated marginal for for that configuration. The output of this integral is the posterior marginal . In practice, we use a discrete set of integration points for , and corresponding weights , to get for which we require and . Usually, the integration is done in a standardised scale, i.e. with respect to . Here, is the mode of and the matrix is the negative square root of the approximated covariance matrix for at the mode.
The relevant options are
opts = control.inla(int.strategy = "user", int.design = Design)where Design is a matrix with the integration points and
the integration weights. The
th
row of Design consists of the values
,
and the integration weight for this configuration,
.
The values are in the
-scale,
meaning that you have to know exactly what you are doing, including
knowing the ordering of the hyperparameters.
Another version, is to define the points in the standardised scale . To do this, use
opts = control.inla(int.strategy = "user.std", int.design = Design)instead. The meaning of Design is unchanged, except that
these can be given in standardised coordinates. This version is more
relevant if you want to implement a generic new integration design
instead of the ones already provided.
First example
In this artificial example, we want to compute the change of the marginal variance of one component, , of a hidden AR process, with respect to lag one correlation . So we want to compute for a fixed value of . We have to compute a numerical approximation, using finite difference. While doing this, it is a good idea to keep the design fixed, to avoid introducing an error for changing that part as well.
Let us first setup the experiment
this gives the following
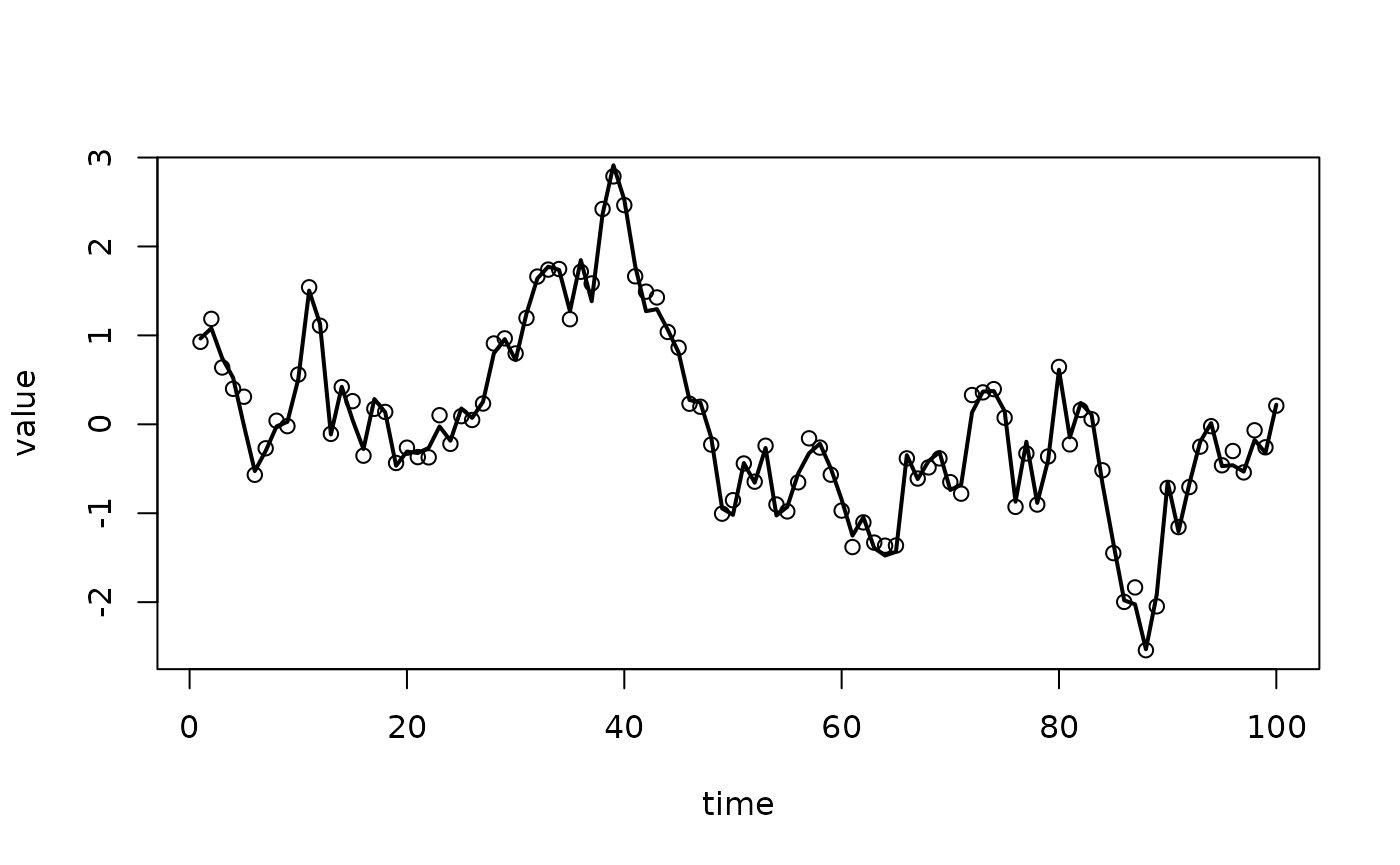 To compute the derivative, we do
To compute the derivative, we do
rho.0 = rho
to.theta = inla.models()$latent$ar1$hyper$theta2$to.theta
rho.0.internal = to.theta(rho.0)
r = inla(y ~ -1 + f(time, model="ar1",
hyper = list(
theta1 = list(prior = "loggamma",
param = c(1,1)),
theta2 = list(initial = rho.0.internal,
fixed=TRUE))),
control.inla = list(int.strategy = "grid"),
data = data.frame(y, time = 1:n))
sd.0 = r$summary.random$time[1,"sd"]
print(sd.0)## [1] 0.01215581We will now change a little, while we keep the same integration points,
summary(r)## Time used:
## Pre = 0.28, Running = 0.176, Post = 0.0173, Total = 0.473
## Random effects:
## Name Model
## time AR1 model
##
## Model hyperparameters:
## mean sd 0.025quant 0.5quant
## Precision for the Gaussian observations 2.20e+04 2.42e+04 1463.817 1.44e+04
## Precision for time 8.36e-01 1.17e-01 0.626 8.29e-01
## 0.975quant mode
## Precision for the Gaussian observations 86134.22 3989.083
## Precision for time 1.09 0.817
##
## Marginal log-Likelihood: -71.09
## is computed
## Posterior summaries for the linear predictor and the fitted values are computed
## (Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')
nm <- nrow(r$summary.hyperpar)
Design = as.matrix(cbind(r$joint.hyper[, seq_len(nm)], 1))
head(Design)## Log precision for the Gaussian observations Log precision for time 1
## [1,] 5.459082 -0.46095952 1
## [2,] 5.459158 -0.29206150 1
## [3,] 5.459209 -0.17946281 1
## [4,] 5.459256 -0.07407056 1
## [5,] 5.459328 0.08401781 1
## [6,] 7.433883 -0.46184951 1where the last column is the (un-normalised) integration weights. The integration weights should depend on the area associated with every point, but for simplicity we just set the weights to be a constant.
Design has dimension 32, 3. We call inla()
again reusing the previous found mode
h.rho = 0.01
rho.1.internal = to.theta(rho.0 + h.rho)
rr = inla(y ~ -1 + f(time, model="ar1",
hyper = list(
theta1 = list(prior = "loggamma",
param = c(1,1)),
theta2 = list(initial = rho.1.internal,
fixed=TRUE))),
control.mode = list(result = r, restart=FALSE),
data = data.frame(y, time = 1:n),
control.inla = list(
int.strategy = "user",
int.design = Design))
sd.1 = rr$summary.random$time[1,"sd"]
print(sd.1)## [1] 0.01045805and then our estimate of the derivative is
deriv.1 = (sd.1 - sd.0) / h.rho
print(deriv.1)## [1] -0.1697763PS: In the logfile of the inla()-call, the
configurations are shown in the
scale even for int.strategy="user".
Second example
There is also another (experimental) option, that is
control.inla = list(int.stategy = "user.expert")for which the weights . The following example show how to use it, fitting the same model in three different ways.
n = 50
x = rnorm(n)
y = 1 + x + rnorm(n, sd = 0.2)
param = c(1, 0.04)
dz = 0.1
r.std = inla(y ~ 1 + x, data = data.frame(y, x),
control.inla = list(int.strategy = "grid",
dz = dz,
diff.logdens = 8),
control.family = list(
hyper = list(
prec = list(
prior = "loggamma",
param = param))))
s = r.std$internal.summary.hyperpar[1,"sd"]
m = r.std$internal.summary.hyperpar[1,"mean"]
theta = m + s*seq(-4, 4, by = dz)
weight = dnorm(theta, mean = m, sd = s)
r = rep(list(list()), length(theta))
for(k in seq_along(r)) {
r[[k]] = inla(y ~ 1 + x,
control.family = list(
hyper = list(
prec = list(
initial = theta[k],
fixed=TRUE))),
data = data.frame(y, x))
}
r.merge = inla.merge(r, prob = weight)## Warning in inla.merge(r, prob = weight): This function is experimental.## Warning in inla.merge(r, prob = weight): Merging 'cpo' and 'pit'-results
## are/can be, approximate only
r.design = inla(y ~ 1 + x,
data = data.frame(y, x),
control.family = list(
hyper = list(
prec = list(
## the prior here does not really matter, as we will override
## it with the user.expert in any case.
prior = "pc.prec",
param = c(1, 0.01)))),
control.inla = list(int.strategy = "user.expert",
int.design = cbind(theta, weight)))
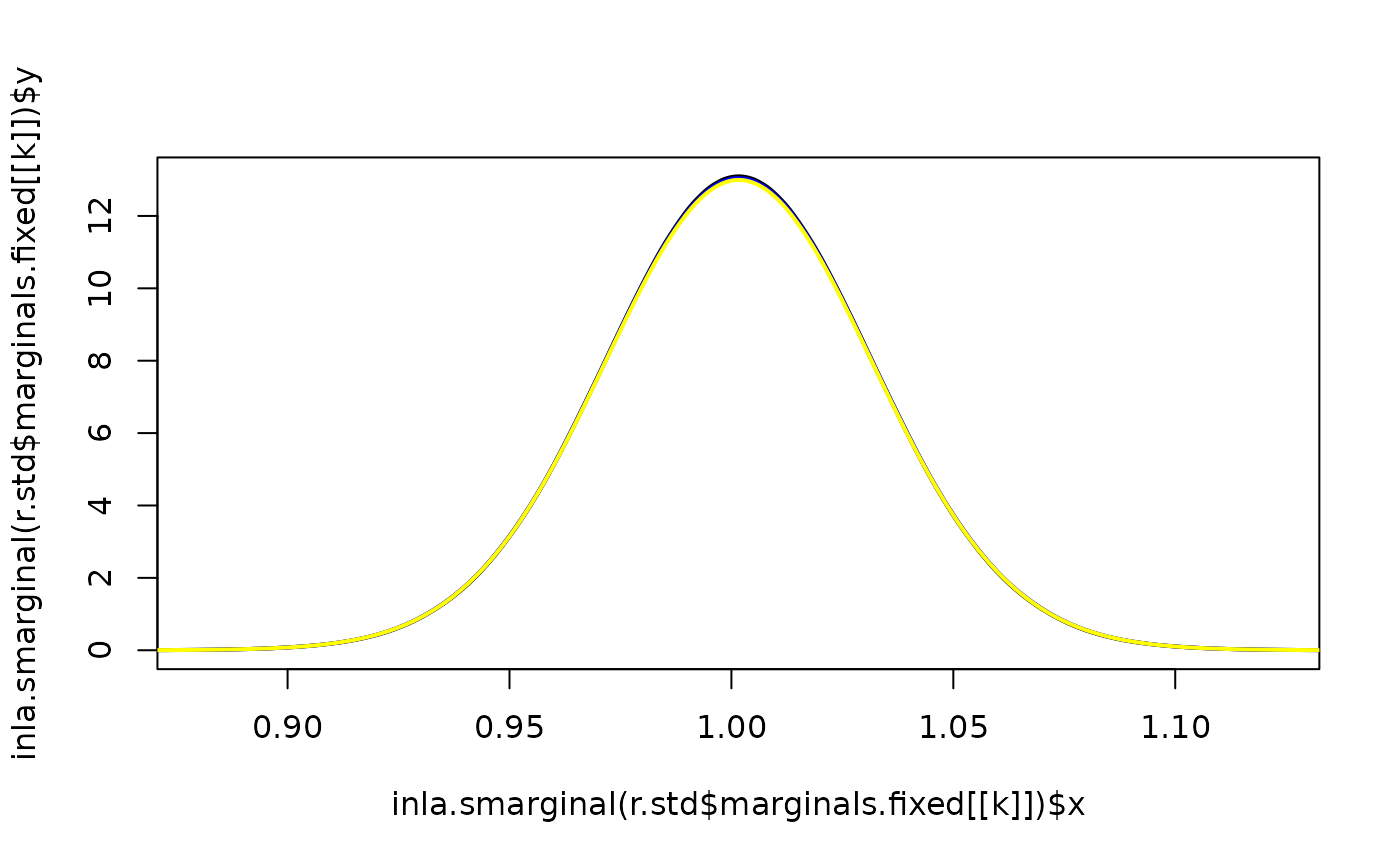
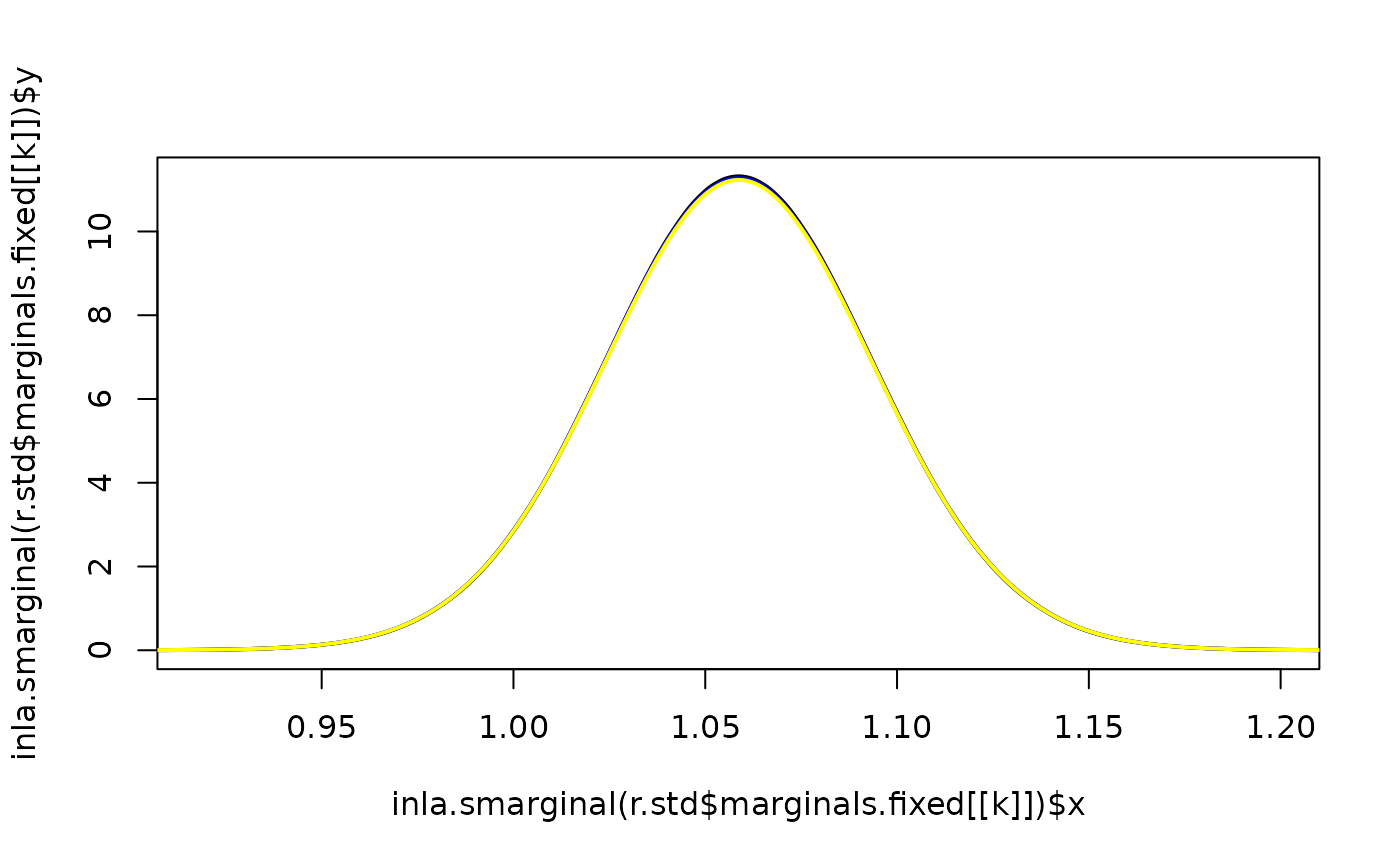
for(k in 1:2) {
plot(inla.smarginal(r.std$marginals.fixed[[k]]),
lwd = 2, lty = 1, type = "l",
xlim = inla.qmarginal(c(0.0001, 0.9999), r.std$marginals.fixed[[k]]))
lines(inla.smarginal(r.design$marginals.fixed[[k]]),
lwd = 2, col = "blue", lty = 1)
lines(inla.smarginal(r.merge$marginals.fixed[[k]]),
lwd = 2, col = "yellow", lty = 1)
}