Contents
- Contents
- Introduction
- User-defined priors for the hyperparameters
- Does INLA support the use of different link-functions?
- How can I do predictions using INLA?
- Some of the models needs a graph, how do I specify it?
- How deal with
- Can INLA deal with missing covariates?
- Compute cross-validation or predictive measures of fit
- I have access to a remote Linux/MacOS server, is it possible to run the computations remotely and running R locally?
- Posteriors for linear combinations
- INLA seems to work great for near all cases, but are there cases where INLA is known to have problems?
- Can I have the linear predictor from one model as a covariate in a different model?
- Latent models, likelihoods and priors.
- Copying a model
- Replicate a model
- Models with more than one type of likelihood
- Models where the response/data depends on linear combinations of the “linear predictor” (or the sum of “fixed” and “random” effects)
Introduction
This vignette is a copy and slightly edited FAQ-entries and other short tutorials from the old page. The content comes is slightly random order, sorry about that. We run all examples here in classic-mode, for this reason.
inla.setOption(inla.mode="classic")User-defined priors for the hyperparameters
If you want to use a prior for the hyperparameter that is not yet implemented there are two choices. If you think that your prior should be on the list and that other might use it to, please let us know. Alternatively, you can define your own prior using \verb|prior = “expression: ….”|, or by specifiying a table of and values which define the prior distribution.
There are three ways to specify prior distributions for hyperparameters in INLA:
- Use an available prior distribution
- Define your own prior distribution function using -like (not equal) syntax as expression.
- Create a table of values which represent your prior distribution.
In the following we will provide more details regarding the two last options. Finally, we will present an example illustrating (and comparing) the three different possibilities by means of the log-gamma distribution for the precision parameter.
A user can specify any (univariate) prior distribution for the hyperparameter by defining an expression for the log-density log , as a function of the corresponding . It is important to be aware that is on the internal scale.
The format is
expression: statement; statement; ...; return(value)where “statement” is any regular statement (more below) and “value” is the value for the log-density of the prior, evaluated at the current value for .
Here, is an example defining the log-gamma distribution:
prior.expression = "expression:
a = 1;
b = 0.1;
precision = exp(log_precision);
logdens = log(b^a) - lgamma(a)
+ (a-1)*log_precision - b*precision;
log_jacobian = log_precision;
return(logdens + log_jacobian);"Some syntax specific notes: * No white-space before “(.)” in the return statement. * A “;” is needed to terminate each expression. * A “_” is allowed in variable names.
Known functions that can be used within the expression statement are
- common math functions, such as exp, sin, …
- “gamma” denotes the gamma-function and “lgamma” is its log
- is expressed as either or
Instead of defining a prior distribution function, it is possible to provide a table of suitable values (internal scale) and the corresponding log-density values . INLA fits a spline through the provided points and continues with this in the succeeding computations. Note, there is no transformation into a functional form performed or required. The input-format for the table is a string, which starts with and is then followed by a block of -values and a block of the corresponding -values, which represent the values of the log-density evaluated on x. Thus
We illustrate all three different ways of defining a prior distribution for the precision of a normal likelihood. To show that the three definitions lead to the same result we inspect the logmarginal likelihood.
## the loggamma-prior
prior.function = function(log_precision) {
a = 1;
b = 0.1;
precision = exp(log_precision);
logdens = log(b^a) - lgamma(a) + (a-1)*log_precision - b*precision;
log_jacobian = log_precision;
return(logdens + log_jacobian)
}
## implementing the loggamma-prior using "expression:"
prior.expression = "expression:
a = 1;
b = 0.1;
precision = exp(log_precision);
logdens = log(b^a) - lgamma(a)
+ (a-1)*log_precision - b*precision;
log_jacobian = log_precision;
return(logdens + log_jacobian);"
## use suitable support points x
lprec = seq(-10, 10, len=100)
## link the x and corresponding y values into a
## string which begins with "table:""
prior.table = paste(c("table:", cbind(lprec,
prior.function(lprec))), collapse=" ", sep="")
# simulate some data
n = 50
y = rnorm(n)
## use the built-in loggamma prior
r1 = inla(y~1,data = data.frame(y),
control.family = list(hyper = list(prec = list(
prior = "loggamma", param = c(1, 0.1)))))
## use the definition using expression
r2 = inla(y~1, data = data.frame(y),
control.family = list(hyper = list(
prec = list(prior = prior.expression))))
## use a table of x and y values representing the loggamma prior
r3 = inla(y~1, data = data.frame(y),
control.family = list(hyper = list(
prec = list(prior = prior.table))))
print(round(c(r1$mlik[1], r2$mlik[1], r3$mlik[1]), digits=3))## [1] -70.657 -70.657 -70.657Does INLA support the use of different link-functions?
Yes, the type of link function is given in the statement using , and the type of link-functions implemented are listed on the documentation for each likelihood. The default link is which corresponds to the second link function in the list. Here is an example
n = 100
z = rnorm(n)
eta = 1 + 0.1*z
N = 2
p = inla.link.invlogit(eta)
y = rbinom(n, size = N, prob = p)
r = inla(y ~ 1 + z, data = data.frame(y, z), family = "binomial", Ntrials = rep(N, n),
control.family = list(control.link = list(model="logit")),
control.predictor = list(compute=TRUE))
p = inla.link.invprobit(eta)
y = rbinom(n, size = N, prob = p)
rr = inla(y ~ 1 + z, data = data.frame(y, z), family = "binomial", Ntrials = rep(N, n),
control.family = list(control.link = list(model="probit")),
control.predictor = list(compute=TRUE))
p = inla.link.invcloglog(eta)
y = rbinom(n, size = N, prob = p)
rrr = inla(y ~ 1 + z, data = data.frame(y, z), family = "binomial", Ntrials = rep(N, n),
control.family = list(control.link = list(model="cloglog")),
control.predictor = list(compute=TRUE)) Other linkfunctions/models are also avilable from within , see
How can I do predictions using INLA?
In there is no function as for in . Predictions must to done as a part of the model fitting itself. As prediction is the same as fitting a model with some missing data, we can simply set for those “locations” we want to predict. Here is a simple example
n = 100
n.pred = 10
y = arima.sim(n=n, model=list(ar=0.9))
N = n + n.pred
yy = c(y, rep(NA, n.pred))
i = 1:N
formula = yy ~ f(i, model="ar1")
r = inla(formula, data = data.frame(i,yy),
control.family = list(initial = 10, fixed=TRUE)) ## no observational noisewhich gives predictions
r$summary.random$i[(n+1):N, c("mean", "sd") ]## mean sd
## 101 1.4373389 1.086169
## 102 1.2311683 1.298859
## 103 1.0584224 1.435747
## 104 0.9131657 1.529731
## 105 0.7905775 1.596595
## 106 0.6867469 1.645303
## 107 0.5984943 1.681415
## 108 0.5232261 1.708570
## 109 0.4588186 1.729236
## 110 0.4035274 1.745132Quantiles such like and , if computed, use the identity link if by default. If you want the computed with a different link function, then there are two ways to doit.
In the case you want to use the link-function from the likelihood already used (most often the case), there is the argument in . If the response , then set , to indicate that you want to compute that fitted value using the link function from . With several likelihoods, set to family-index which is correct, ie the column number in the response. The following example shows the usage:
## simple poisson regression
n = 100
x = sort(runif(n))
eta = 1 + x
lambda = exp(eta)
y = rpois(n, lambda = lambda)
## missing values:
y[1:3] = NA
y[(n-2):n] = NA
## link = 1 is a shortcut for rep(1, n) where n is the appropriate
## length. here '1' is a reference to the first 'family', ie
## 'family[1]'
r = inla(y ~ 1 + x, family = "poisson",
data = data.frame(y, x),
control.predictor = list(link = 1))
plot(exp(eta),type ="l")
points(r$summary.fitted.values$mean, pch=19)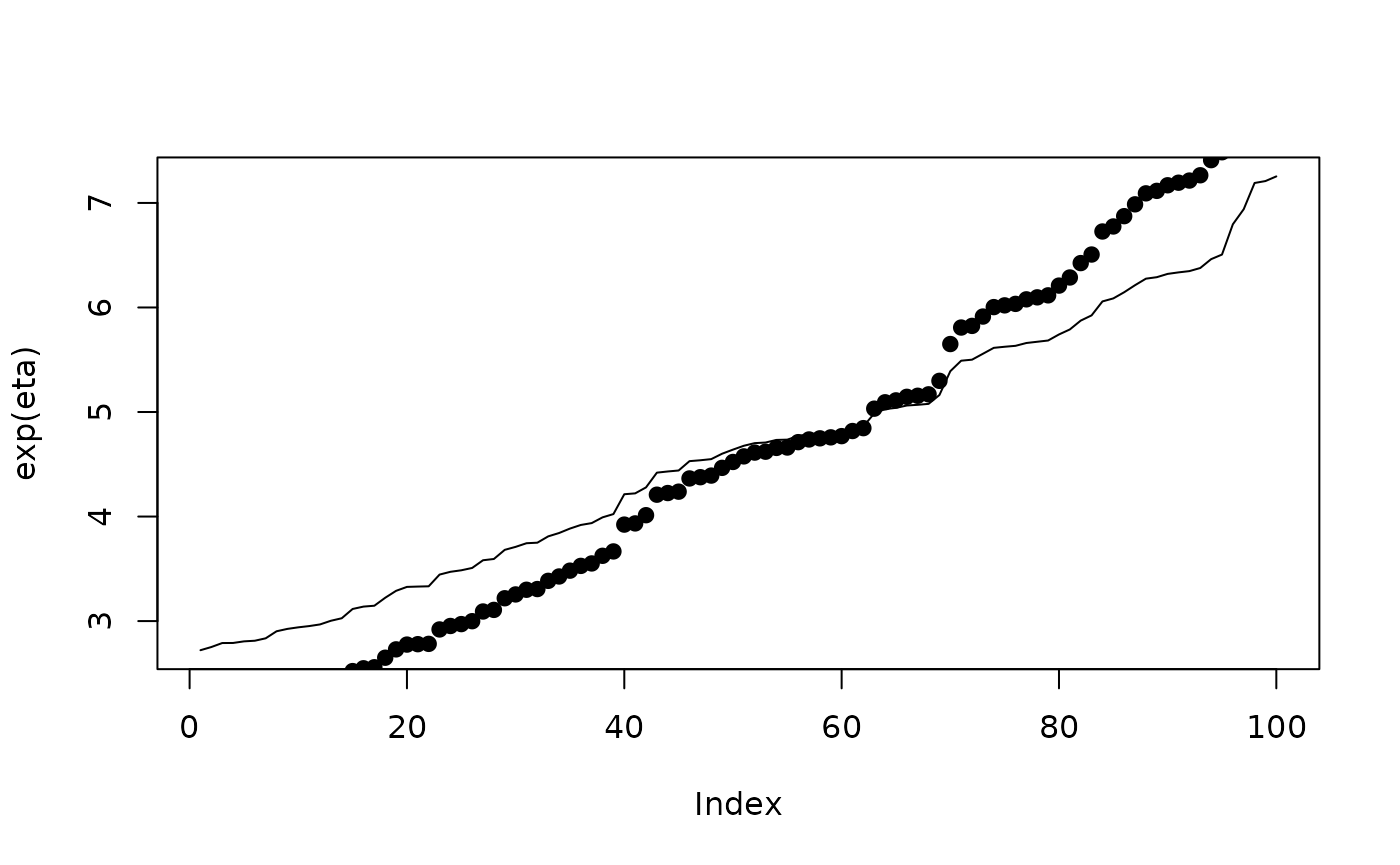
We only need to define where there are missing values. Entries for which the observation is not , is ignored.
For more than one likelihood, use ‘2’ to refer to the second likelihood. Here is an example where we split the data in two, and assign the second half the distribution.
n2 = n %/% 2L
Y = matrix(NA, n, 2)
Y[1:n2, 1] = y[1:n2]
Y[1:n2 + n2, 2] = y[1:n2 + n2]
link = rep(NA, n)
link[which(is.na(y[1:n2]))] = 1
link[n2 + which(is.na(y[1:n2 + n2]))] = 2
r = inla(Y ~ 1 + x, family = c("poisson", "nbinomial"),
data = list(Y=Y, x=x),
control.family= list(list(), list()),
control.predictor = list(link = link))
plot(exp(eta),type ="l")
points(r$summary.fitted.values$mean, pch=19)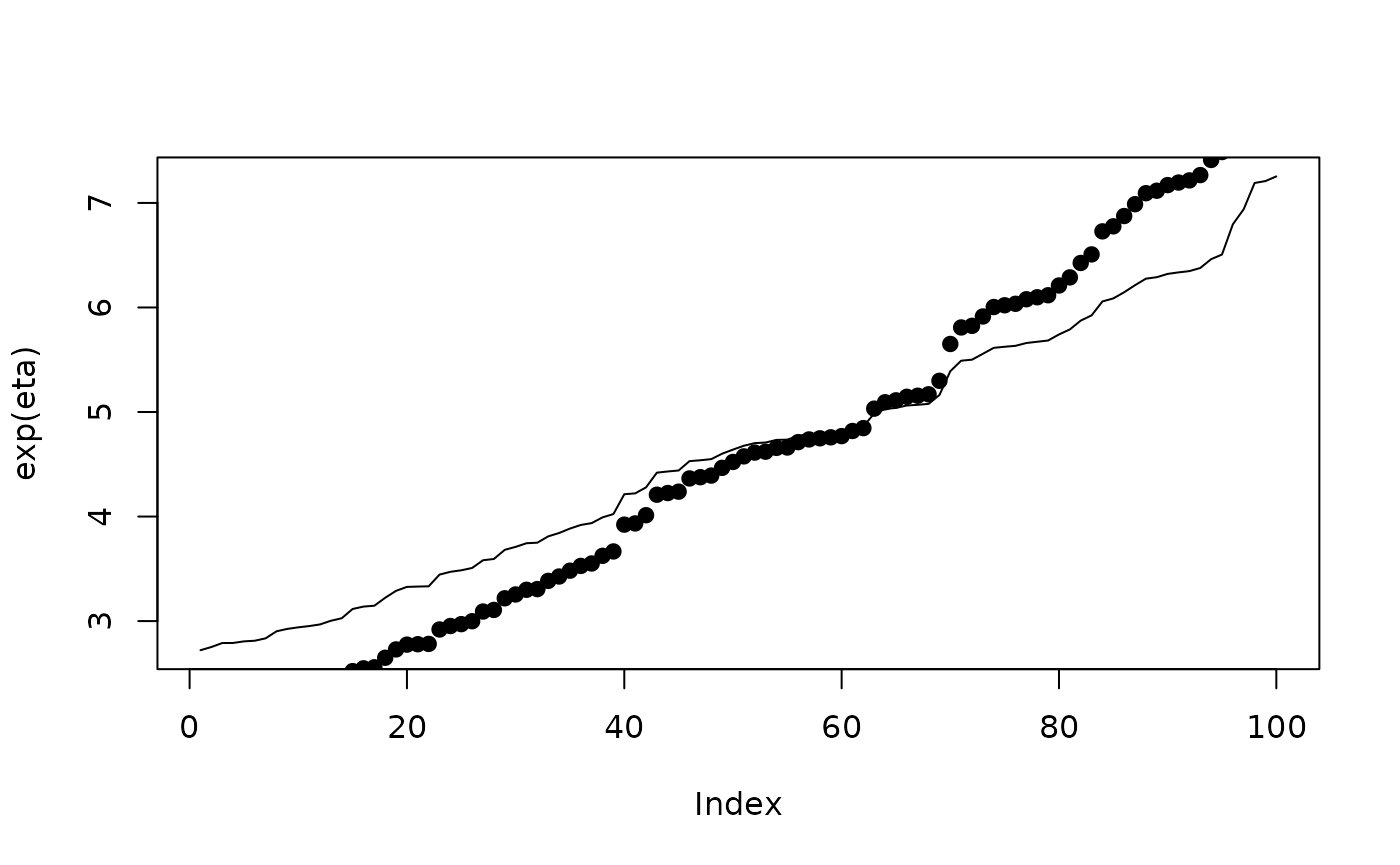
We can transform marginals manually using the function or compute expectations using , like in this example (taken from ).
## y n time
## Min. :0.0000 Min. :1.000 Min. : 1.00
## 1st Qu.:0.0000 1st Qu.:2.000 1st Qu.: 92.25
## Median :0.0000 Median :2.000 Median :183.50
## Mean :0.5246 Mean :1.997 Mean :183.50
## 3rd Qu.:1.0000 3rd Qu.:2.000 3rd Qu.:274.75
## Max. :2.0000 Max. :2.000 Max. :366.00
Tokyo$y[300:366] <- NA
## Define the model
formula = y ~ f(time, model="rw2", scale.model=TRUE,
constr=FALSE, cyclic=TRUE,
hyper = list(prec=list(prior="pc.prec",
param=c(2,0.01)))) -1
## We'll get a warning since we have not defined the link argument
result = inla(formula, family="binomial", Ntrials=n, data=Tokyo,
control.compute = list(return.marginals.predictor = TRUE),
control.predictor=list(compute=T))
## need to recompute the fitted values for those with data[i] = NA,
## as the identity link is used.
n = 366
fitted.values.mean = numeric(n)
for(i in 1:366) {
if (is.na(Tokyo$y[i])) {
if (FALSE) {
## either like this, which is slower
marg = inla.marginal.transform(
function(x) exp(x)/(1+exp(x)),
result$marginals.fitted.values[[i]] )
fitted.values.mean[i] = inla.emarginal(function(x) x, marg)
} else {
## or like this, which is faster
fitted.values.mean[i] = inla.emarginal(
function(x) exp(x)/(1 +exp(x)),
result$marginals.fitted.values[[i]])
}
} else {
fitted.values.mean[i] = result$summary.fitted.values[i,"mean"]
}
}
plot(fitted.values.mean)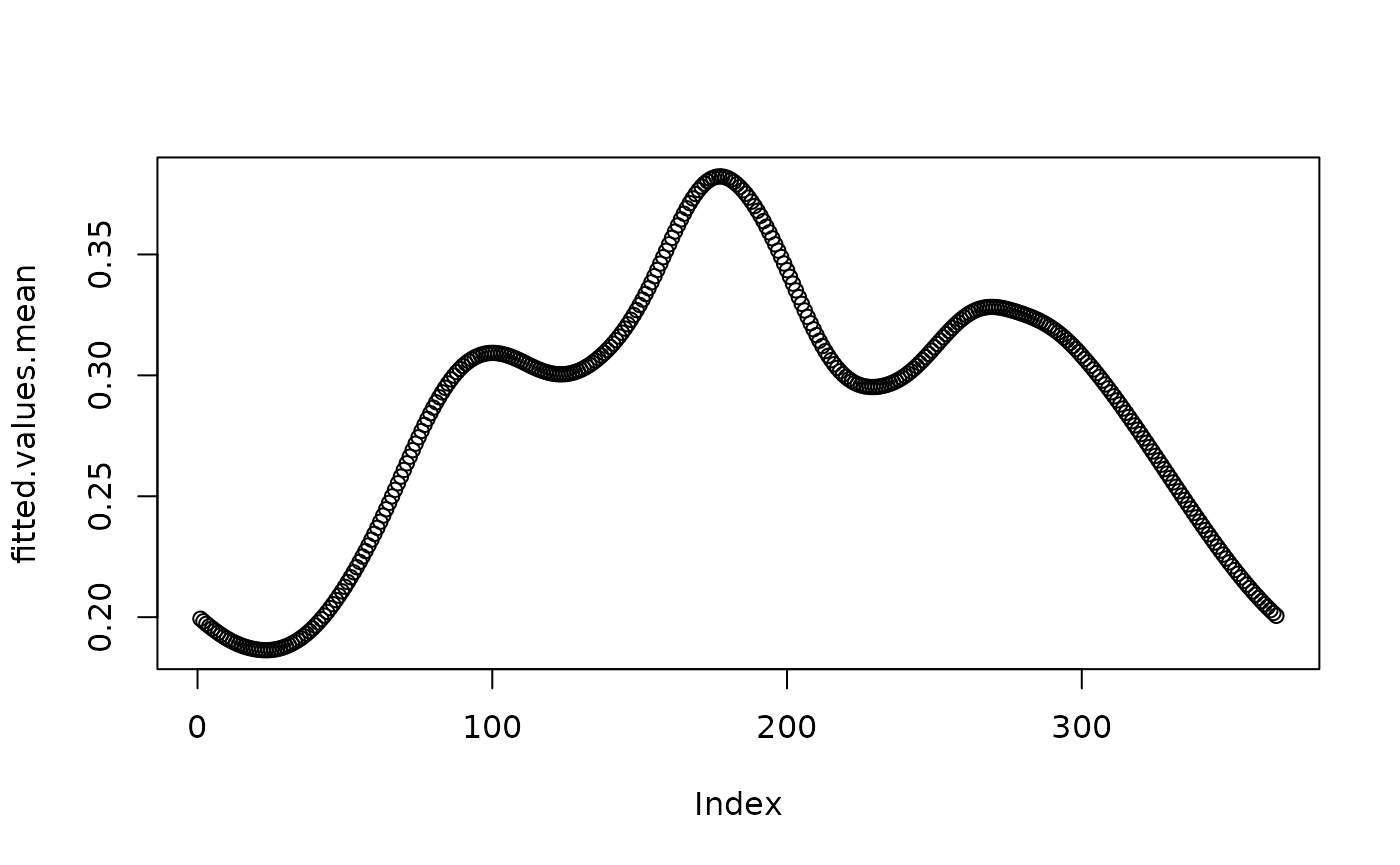
Some of the models needs a graph, how do I specify it?
Some of the models in needs the user to specify a graph, saying which nodes are neighbours to each other. A ‘graph’ can be specified in three different ways.
- As an ascii or binary file with a graph specification, or the same contents given as (possible list of) mix of character and numerics arguments.
- As a symmetric (dense or sparse) matrix, where the non-zero pattern of the matrix defines the graph.
- As an inla.graph-object
A graph defined in an ascii-file, must have the following format. The first entry is the number of nodes in the graph, . The nodes in the graph are labelled . The next entries, specify the number of neighbours and the neighbours for each node. A simple example is the following
This defines a graph with four nodes, where node 1 has 2 neighbours 3
and 4, node 2 as 0 neighbours, node 3 has 1 neighbour 1, and node 4 has
1 neighbour 1, and the graph looks like this (requires
Rgraphviz and graph)
g = inla.read.graph("4 1 2 3 4 2 0 3 1 1 4 1 1")
if (FALSE && requireNamespace("Rgraphviz") && requireNamespace("graph")) {
tryCatch(
plot(g),
error = function(e) {
message("Rgraphviz plotting doesn't seem to work on this installation.")
}
)
}Note that we need to specify the implied symmetry as well. In this example 4 is a neighbour of 1, then we also need to specify that 1 is a neighbour of 4.
Instead of storing the graph specification on a file, it can also be specified as a character string with the same contents as a file, like
"4 1 2 3 4 2 0 3 1 1 4 1 1"as used in above.
Due to imitations of the length of a string/line, so in practice, this way specifying the graph, seems more useful for teaching or demonstration purposes than for practical analysis.
Within , this would look like
formula = y ~ f(idx, model = "besag", graph = "graph.dat")or
formula = y ~ f(idx, model = "besag", graph = "4 1 2 3 4 2 0 3 1 1 4 1 1")A graph can also be defined as a symmetric (dense or sparse) matrix, where the non-zero pattern of the matrix defines the graph A neighbour matrix is often used for defining which nodes that are neighbours, with the convention that if then and are neighbours if .
For example, the (dense) matrix C
## [,1] [,2] [,3] [,4]
## [1,] 1 0 1 1
## [2,] 0 1 0 0
## [3,] 1 0 1 0
## [4,] 1 0 0 1defines the same graph show above.
Since graphs tends to be large, we often define them as a sparse matrix
C.sparse= inla.as.sparse(C)
C.sparse## 4 x 4 sparse Matrix of class "dgTMatrix"
##
## [1,] 1 . 1 1
## [2,] . 1 . .
## [3,] 1 . 1 .
## [4,] 1 . . 1We can then use or in the formula.
We can also define the graph as an -object, which is used internally, represent a graph. For example
str(g)## List of 4
## $ n : int 4
## $ nnbs: num [1:4] 2 0 1 1
## $ nbs :List of 4
## ..$ : int [1:2] 3 4
## ..$ : num(0)
## ..$ : int 1
## ..$ : int 1
## $ cc :List of 4
## ..$ id : int [1:4] 1 2 1 1
## ..$ n : int 2
## ..$ nodes:List of 2
## .. ..$ : int [1:3] 1 3 4
## .. ..$ : int 2
## ..$ mean : Factor w/ 1 level "1": 1 NA 1 1
## - attr(*, "class")= chr "inla.graph"and use as the argument.
The internal format, are as follows. is the size of the graph. are the number of neighbours to each node, list all the neighbours to each node, and the class is . The -list is for internal use only and specify the connected components in the graph.
has some functions to work with graphs, and here is a short summary.
and , read and write graphs using any of the graph specifications above.
You can plot a -object using and get a summary using . The plotting requires the package.
From a graph specification, you can generate the neighbour matrix, using . You can plot a graph specification as a neighbour matrix, using
If you have ‘errors’ in your graph, you may read it using . This is only available for a graph specification in an ascii-file.
How deal with
For a formula like
then ’s in either , or are treated differently.
’s in the response . If , this means that is not observed, hence gives no contribution to the likelihood.
’s in fixed effect . If this means that is not part of the linear predictor for . For fixed effects, this is equivalent to , hence internally we make this change: .
’s in random effect . If , this means that the random effect does not contribute to the linear predictor for .
’s in a factor . ’s in a factor is not allowed unless is a level in itself, or
control.fixed = list(expand.factor.strategy = "inla")is set. With this option, then is interpreted similarly as a fixed effect, where means no contribution from x. The effect of , is best explained with an example.
r = inla(y ~ 1 + x, data = data.frame(y=1:3, x=factor(c("a","b","c"))))
as.matrix(r$model.matrix)## (Intercept) xb xc
## 1 1 0 0
## 2 1 1 0
## 3 1 0 1for default value of the argument . The effect of is removed to make the corresponding matrix non-singular. If we want to expand into each of each three effects, then we can do
r = inla(y ~ 1 + x, data = data.frame(y=1:3,x=factor(c("a","b","c"))),
control.fixed = list(expand.factor.strategy="inla"))
as.matrix(r$model.matrix)## (Intercept) xa xb xc
## 1 1 1 0 0
## 2 1 0 1 0
## 3 1 0 0 1As we see, each level of the factor is now treated symetrically. Although the corresponding frequentist matrix is singular as we have confounding with the intercept, the Bayesian posterior is still proper with proper priors.
With a in , we get
r = inla(y ~ 1 + x, data = data.frame(y=1:3,x=factor(c("a","b",NA))),
control.fixed = list(expand.factor.strategy="inla"))
as.matrix(r$model.matrix)## (Intercept) xa xb
## 1 1 1 0
## 2 1 0 1
## 3 1 0 0so that the 3rd element of the linear predictor has no contribution from , as it should.
Can INLA deal with missing covariates?
No, INLA has no generic way to “impute” or integrate-out missing covariates. You have to adjust your model to account for missing covariates, like using one of the measurement error models (“meb”, “mec”), or construct a joint model for the data and the covariates, but this is case-specific.
Compute cross-validation or predictive measures of fit
provides two types of leave-one-out predictive measures of fit. It
is the CPO value, which is
the PIT value
To enable the computation of these
quantities, you will need to add the argument
control.compute=list(cpo=TRUE)We can also compute PO values
when argument is added.
If the resulting object is , then you will find the predictive quantities as and .
Implicit assumptions made in for computations, and there are internal checks that these are satisfied. The results of these checks will appear as . In short, if then some assumption is violated, the higher the value (maximum 1) the more seriously. If then the assumptions should be ok.
You want to recompute those with non-zero failure. However, this must be done by removing from the dataset, fit the model and then predict . To provide a more efficient implementation of this, we have provided
improved.result = inla.cpo(result)which take an inla-object which is the output from , and recompute (in an efficient way) the cpo/pit for which , and return ‘result’ with the improved estimates of cpo/pit. See for details.
I have access to a remote Linux/MacOS server, is it possible to run the computations remotely and running R locally?
Yes! This option allow to use a remote server to do the computations. In order to use this feature, you need to do some setup which is different from (various) Linux distributions, Mac and Windows. In short:
- install and a remote server, for example .
- Install your public ssh-key on to setup password free access to the remove server using . And please check that this is indeed working before moving forward!
- On your local host, run to initialise the init-file and then edit this file to fit your needs.
- You may now have to log out and log in again, to make sure your ssh key is signed out.
- You can now use option to do the computations on your remote server, or set this globally with
You can also submit a job on the remote server, so you do not need to sit and wait for it to finish, but you can collect the results later. Basically, you do
r = inla(..., inla.call = "submit")which will start the job on the server. You can start many jobs, and list them using
and you can fetch the results (for the job above) using
r = inla.qget(r)You can also delete jobs and fetch the jobs from another machine; see for further details.
HOWTO setup ssh-keys: For the unexperienced user, this is somewhat tricky; sorry about that. The easiest is to find a friend that knows this and can help you. Newer system do a lot of these things very nicely these days.
It is also possible to setup this from Windows using CYGWIN, and INLA can work with this interface as well. Please see the old web-page for details, which are long and technical. HOWEVER, I am no longer convinced that this work anymore, as I haven’t seen this is use for years. It is much much easier to use a virtual machine with Linux on Windows.
Posteriors for linear combinations
I have some linear combinations of the nodes in the latent field that I want to compute the posterior marginal of, is that possible? Yes! These are called ‘linear combinations’. There are handy functions, ‘inla.make.lincomb()’ and ‘inla.make.lincombs()’, to define one or many such linear combinations. Single linear combinations made by using ‘inla.make.lincomb()’ can easily be joined into many. Its use is easiest explained using a rather long example…
Here is the example, that explains these features.
## A simple model
n = 100
a = rnorm(n)
b = rnorm(n)
idx = 1:n
y = rnorm(n) + a + b
formula = y ~ 1 + a + b + f(idx, model="iid")
## assume we want to compute the posterior for
##
## 2 * beta_a + 3 * beta_b + idx[1] - idx[2]
##
## which we specify as follows (and giving it a unique name)
lc1 = inla.make.lincomb(a=2, b=3, idx = c(1,-1,rep(NA,n-2)))
names(lc1) = "lc1"
## strictly speaking, it is sufficient to use `idx = c(1,-1)', as the
## remaining idx's are not used in any case.
r = inla(formula, data = data.frame(a,b,y),
## add the linear combinations here
lincomb = lc1,
## force noise variance to be essiatially zero
control.family = list(initial=10, fixed=TRUE))
## to verify the result, we can compare the mean but the variance and
## marginal cannot be computed from the simpler marginals alone.
lc1.1 = 2 * r$summary.fixed["a", "mean"] + 3 * r$summary.fixed["b",
"mean"] + r$summary.random$idx$mean[1] -
r$summary.random$idx$mean[2]
lc1.2= r$summary.lincomb.derived$mean
print(round(c(lc1.1 = lc1.1, lc1.2 = lc1.2), dig=3))## lc1.1 lc1.2
## 6.904 6.904The marginals are available as
There is an another function which is handy for specifying many linear combinations at once, that is (note the plural s). Here each ‘row’ define one linear combination
## let wa and wb be vectors, and we want to compute the marginals for
## beta_a * wa[i] + beta_b * wb[i], for i=1..m. this is done
## conveniently as follows
m = 10
wa = runif(m)
wb = runif(m)
lc.many = inla.make.lincombs(a = wa, b=wb)
## we can give them names as well, but there are also default names, like
print(names(lc.many))## [1] "lc01" "lc02" "lc03" "lc04" "lc05" "lc06" "lc07" "lc08" "lc09" "lc10"
r = inla(formula, data = data.frame(a,b,y),
lincomb = lc.many,
control.family = list(initial=10, fixed=TRUE))
print(round(r$summary.lincomb.derived, dig=3))## ID mean sd 0.025quant 0.5quant 0.975quant mode kld
## lc01 1 0.380 0.037 0.306 0.380 0.453 0.380 0
## lc02 2 0.980 0.080 0.823 0.980 1.137 0.980 0
## lc03 3 1.024 0.087 0.854 1.024 1.194 1.024 0
## lc04 4 1.106 0.078 0.952 1.106 1.260 1.106 0
## lc05 5 1.580 0.115 1.354 1.580 1.807 1.580 0
## lc06 6 1.287 0.088 1.115 1.287 1.460 1.287 0
## lc07 7 0.504 0.034 0.437 0.504 0.571 0.504 0
## lc08 8 1.571 0.108 1.359 1.571 1.783 1.571 0
## lc09 9 0.978 0.066 0.848 0.978 1.108 0.978 0
## lc10 10 0.575 0.039 0.498 0.575 0.652 0.575 0Terms like ‘idx’ above, can be added as into , where IDX is a matrix. Again, each row of the arguments define one linear combination.
There is a further option available for the derived linear combinations, that is the option to compute also the posterior correlation matrix between all the linear combinations. To activate this option, use
control.inla = list(lincomb.derived.correlation.matrix = TRUE)and you will find the resulting posterior correlation matrix as
result$misc$lincomb.derived.correlation.matrixHere is a small example where we compute the correlation matrix for the predicted values of a hidden AR(1) model with an intercept.
n = 100
nPred = 10
phi = 0.9
x = arima.sim(n, model = list(ar=phi)) * sqrt(1-phi^2)
y = 1 + x + rnorm(n, sd=0.1)
time = 1:(n + nPred)
Y = c(y, rep(NA, nPred))
formula = Y ~ 1 + f(time, model="ar1")
## make linear combinations which are the nPred linear predictors
B = matrix(NA, nPred, n+nPred)
for(i in 1:nPred) {
B[i, n+i] = 1
}
lcs = inla.make.lincombs(Predictor = B)
r = inla(formula, data = data.frame(Y, time),
control.predictor = list(compute=TRUE),
lincomb = lcs,
inla.mode="classic",
control.inla = list(lincomb.derived.correlation.matrix=TRUE))
print(round(r$misc$lincomb.derived.correlation.matrix,dig=3))## lc01 lc02 lc03 lc04 lc05 lc06 lc07 lc08 lc09 lc10
## lc01 1.000 0.677 0.529 0.439 0.376 0.329 0.292 0.262 0.237 0.216
## lc02 0.677 1.000 0.781 0.647 0.554 0.484 0.430 0.385 0.348 0.317
## lc03 0.529 0.781 1.000 0.828 0.708 0.619 0.548 0.491 0.444 0.404
## lc04 0.439 0.647 0.828 1.000 0.855 0.746 0.661 0.591 0.534 0.485
## lc05 0.376 0.554 0.708 0.855 1.000 0.872 0.772 0.690 0.622 0.565
## lc06 0.329 0.484 0.619 0.746 0.872 1.000 0.884 0.790 0.712 0.646
## lc07 0.292 0.430 0.548 0.661 0.772 0.884 1.000 0.893 0.804 0.729
## lc08 0.262 0.385 0.491 0.591 0.690 0.790 0.893 1.000 0.899 0.814
## lc09 0.237 0.348 0.444 0.534 0.622 0.712 0.804 0.899 1.000 0.905
## lc10 0.216 0.317 0.404 0.485 0.565 0.646 0.729 0.814 0.905 1.000INLA seems to work great for near all cases, but are there cases where INLA is known to have problems?
The methodology needs the full conditional density for the latent field to be “near” Gaussian. This is usually achived by either replications or smoothing/“borrowing strength”. A simple example which do not have this, is the following:
n = 100
u = rnorm(n)
eta = 1 + u
p = exp(eta)/(1+exp(eta))
y = rbinom(n, size=1, prob = p)
idx = 1:n
result = inla(y ~ 1 + f(idx, model="iid",
hyper = list(prec = list(prior="pc.prec",
prior = c(1,0.01)))),
data =data.frame(y,idx), family = "binomial",
Ntrials = 1)
summary(result)## Time used:
## Pre = 0.127, Running = 0.174, Post = 0.0105, Total = 0.311
## Fixed effects:
## mean sd 0.025quant 0.5quant 0.975quant mode kld
## (Intercept) 0.757 0.215 0.345 0.754 1.188 0.754 0
##
## Random effects:
## Name Model
## idx IID model
##
## Model hyperparameters:
## mean sd 0.025quant 0.5quant 0.975quant mode
## Precision for idx 88096.17 1526249.04 8.94 211.26 144210.90 13.29
##
## Marginal log-Likelihood: -63.39
## is computed
## Posterior summaries for the linear predictor and the fitted values are computed
## (Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')For each binary observation there is an iid “random effect” , and there is no smoothing/``borrowing strength’’ (apart from the weak intercept). If you plot the loglikelihood for eta for , say, then its an increasing function for increasing eta, so the likelihood itself would like . With an unknown precision for we run into problems; INLA has a tendency to estimate a to high precision for . However, it must be noted that the model is almost singular and you’ll have a strong prior sensitivity in the (exact) results as well. There is a similar discussion in here as well for the Salamander data example.
Can I have the linear predictor from one model as a covariate in a different model?
Yes, this is possible. Essentially, you have to set the linear predictor for the first model equal to ‘u’, and then you can copy ‘u’ and use the scaling to get the regression coefficient. A simple example will illustrate the idea:
## simple example
set.seed(12345L)
sd.data = 1
n = 2000
## fixed high and low precisions are specified with parameters
fixed.high <- 10
fixed.low <- - 6
## otherwise is the model identical to the previous FAQ-model
x1 = rnorm(n)
eta1 = 1 + x1
x2 = rnorm(n)
eta2 = 2 + 2*eta1 + 2*x2
y1 = rnorm(n, mean=eta1, sd = sd.data)
y2 = rnorm(n, mean=eta2, sd = sd.data)
## the trick is to create a vector 'u' (iid) which is equal to eta1, and then we can copy 'u' to
## create beta*u or beta*eta1. we do this by using 0 = eta1 -u + tiny.noise
formula = Y ~ -1 + intercept1 + X1 + intercept2 +
f(u, w, model="iid", hyper = list(prec = list(initial = fixed.low, fixed=TRUE))) +
f(b.eta2, copy="u", hyper = list(beta = list(fixed = FALSE))) + X2
Y = matrix(NA, 3*n, 3)
## part 1: y1
intercept1 = rep(1, n)
X1 = x1
intercept2 = rep(NA, n)
u = rep(NA, n)
w = rep(NA, n)
b.eta2 = rep(NA, n)
X2 = rep(NA, n)
Y[1:n, 1] = y1
## part 2: 0 = eta1 - u + tiny.noise
intercept1 = c(intercept1, intercept1)
X1 = c(X1, x1)
intercept2 = c(intercept2, rep(NA, n))
u = c(u, 1:n)
w = c(w, rep(-1, n))
b.eta2 = c(b.eta2, rep(NA, n))
X2 = c(X2, rep(NA, n))
Y[n + 1:n, 2] = 0
## part 3: y2
intercept1 = c(intercept1, rep(NA, n))
X1 = c(X1, rep(NA, n))
intercept2 = c(intercept2, rep(1, n))
u = c(u, rep(NA, n))
w = c(w, rep(NA, n))
b.eta2 = c(b.eta2, 1:n)
X2 = c(X2, x2)
Y[2*n + 1:n, 3] = y2
r = inla(formula,
data = list(Y=Y, intercept1=intercept1, X1=X1,
intercept2=intercept2, u=u, w=w, b.eta2=b.eta2, X2=X2),
family = c("gaussian", "gaussian", "gaussian"),
control.family = list(
list(),
list(hyper = list(prec = list(initial = fixed.high, fixed=TRUE))),
list()))
summary(r)## Time used:
## Pre = 0.165, Running = 2.3, Post = 0.127, Total = 2.59
## Fixed effects:
## mean sd 0.025quant 0.5quant 0.975quant mode kld
## intercept1 0.977 0.023 0.932 0.977 1.021 0.977 0
## X1 1.008 0.022 0.966 1.008 1.050 1.008 0
## intercept2 2.053 0.070 1.915 2.055 2.187 2.055 0
## X2 2.031 0.024 1.984 2.031 2.077 2.031 0
##
## Random effects:
## Name Model
## u IID model
## b.eta2 Copy
##
## Model hyperparameters:
## mean sd 0.025quant 0.5quant
## Precision for the Gaussian observations 0.969 0.031 0.91 0.969
## Precision for the Gaussian observations[3] 0.969 0.031 0.91 0.969
## Beta for b.eta2 1.994 0.049 1.90 1.994
## 0.975quant mode
## Precision for the Gaussian observations 1.03 0.969
## Precision for the Gaussian observations[3] 1.03 0.968
## Beta for b.eta2 2.09 1.992
##
## Marginal log-Likelihood: -13642.63
## is computed
## Posterior summaries for the linear predictor and the fitted values are computed
## (Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')
## An alternative specification of the same model applies the inla.stack() function.
## It is called four times, one time for each submodel, one time for the "copying",
## and one time for joining the three substacks to one.
## Part 1:
## The first model stack comes in two versions
mod1a.stack <- inla.stack(
data = list(Y = as.matrix(cbind(y1, NA, NA))),
A = list(1, 1),
effects = list(intercept1 = rep(1,n),
x1 = x1)
)
mod1b.stack <- inla.stack(
data = list(Y = as.matrix(cbind(y1, NA, NA))),
A = list(1),
effects = list(u = 1:n)
)
## Part 2:
copy.stack <- inla.stack(
data = list(Y = as.matrix(cbind(NA, rep(0,n), NA))),
A = list(1, 1, - 1),
effects = list(intercept1 = rep(1,n),
x1 = x1,
u = 1:n)
)
## Part 3:
mod2.stack <- inla.stack(
data = list(Y = as.matrix(cbind(NA, NA, y2))),
A = list(1, 1, 1),
effects = list(intercept2 = rep(1,n),
x2 = x2,
b.eta2 = 1:n)
)
all.a.stack <- inla.stack(mod1a.stack, copy.stack, mod2.stack)
all.b.stack <- inla.stack(mod1b.stack, copy.stack, mod2.stack)
## The formula is slightly modified because the long vectors X1 and X2 has not been created
## and there is no weight applied as "-1" appears as the effect of u
formula = Y ~ -1 + intercept1 + x1 + intercept2 + x2 +
f(u, model = "iid", hyper = list(prec = list(initial = fixed.low, fixed = TRUE))) +
f(b.eta2, copy = "u", hyper = list(beta = list(fixed = FALSE)))
## The inla() function makes use of the stack
r.new.a = inla(formula,
data = inla.stack.data(all.a.stack),
control.predictor = list(A = inla.stack.A(all.a.stack)),
family = c("gaussian", "gaussian", "gaussian"),
control.family = list(
list(),
list(hyper = list(prec = list(initial = fixed.high, fixed=TRUE))),
list()))
summary(r.new.a)## Time used:
## Pre = 0.186, Running = 3.27, Post = 0.315, Total = 3.77
## Fixed effects:
## mean sd 0.025quant 0.5quant 0.975quant mode kld
## intercept1 0.977 0.023 0.932 0.977 1.021 0.977 0
## x1 1.008 0.022 0.966 1.009 1.050 1.009 0
## intercept2 2.054 0.070 1.915 2.055 2.187 2.056 0
## x2 2.031 0.024 1.984 2.031 2.077 2.031 0
##
## Random effects:
## Name Model
## u IID model
## b.eta2 Copy
##
## Model hyperparameters:
## mean sd 0.025quant 0.5quant
## Precision for the Gaussian observations 0.969 0.031 0.91 0.969
## Precision for the Gaussian observations[3] 0.969 0.031 0.91 0.969
## Beta for b.eta2 1.994 0.049 1.90 1.993
## 0.975quant mode
## Precision for the Gaussian observations 1.03 0.968
## Precision for the Gaussian observations[3] 1.03 0.969
## Beta for b.eta2 2.09 1.990
##
## Marginal log-Likelihood: -13642.62
## is computed
## Posterior summaries for the linear predictor and the fitted values are computed
## (Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')
r.new.b = inla(formula,
data = inla.stack.data(all.b.stack),
control.predictor = list(A = inla.stack.A(all.b.stack)),
family = c("gaussian", "gaussian", "gaussian"),
control.family = list(
list(),
list(hyper = list(prec = list(initial = fixed.high, fixed=TRUE))),
list()))
summary(r.new.b)## Time used:
## Pre = 0.176, Running = 3.21, Post = 0.0839, Total = 3.47
## Fixed effects:
## mean sd 0.025quant 0.5quant 0.975quant mode kld
## intercept1 0.977 0.023 0.932 0.977 1.021 0.977 0
## x1 1.008 0.022 0.966 1.008 1.050 1.008 0
## intercept2 2.053 0.070 1.915 2.055 2.186 2.055 0
## x2 2.031 0.024 1.984 2.031 2.077 2.031 0
##
## Random effects:
## Name Model
## u IID model
## b.eta2 Copy
##
## Model hyperparameters:
## mean sd 0.025quant 0.5quant
## Precision for the Gaussian observations 0.970 0.031 0.91 0.969
## Precision for the Gaussian observations[3] 0.969 0.031 0.91 0.969
## Beta for b.eta2 1.994 0.049 1.90 1.994
## 0.975quant mode
## Precision for the Gaussian observations 1.03 0.969
## Precision for the Gaussian observations[3] 1.03 0.969
## Beta for b.eta2 2.09 1.992
##
## Marginal log-Likelihood: -13642.61
## is computed
## Posterior summaries for the linear predictor and the fitted values are computed
## (Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')
## All specifications give similar and acceptable results both for fixed effects and precisions,
## The version r.new.b seems faster than r.new.a for more complex models with one or more spde-termsLatent models, likelihoods and priors.
The list of latent models, likelihood and priors implemented, can be found by doing (or give a specific section, see )
## Section [group]
## ar AR(p) correlations
## ar1 AR(1) correlations
## besag Besag model
## exchangeable Exchangeable correlations
## exchangeablepos Exchangeable positive correlations
## iid Independent model
## rw1 Random walk of order 1
## rw2 Random walk of order 2
## Section [hazard]
## iid An iid model for the log-hazard
## rw1 A random walk of order 1 for the log-hazard
## rw2 A random walk of order 2 for the log-hazard
## Section [latent]
## 2diid (This model is obsolute)
## ar Auto-regressive model of order p (AR(p))
## ar1 Auto-regressive model of order 1 (AR(1))
## ar1c Auto-regressive model of order 1 w/covariates
## besag The Besag area model (CAR-model)
## besag2 The shared Besag model
## besagproper A proper version of the Besag model
## besagproper2 An alternative proper version of the Besag model
## bym The BYM-model (Besag-York-Mollier model)
## bym2 The BYM-model with the PC priors
## cgeneric Generic latent model specified using C
## clinear Constrained linear effect
## copy Create a copy of a model component
## crw2 Exact solution to the random walk of order 2
## dmatern Dense Matern field
## fgn Fractional Gaussian noise model
## fgn2 Fractional Gaussian noise model (alt 2)
## generic A generic model
## generic0 A generic model (type 0)
## generic1 A generic model (type 1)
## generic2 A generic model (type 2)
## generic3 A generic model (type 3)
## iid Gaussian random effects in dim=1
## iid1d Gaussian random effect in dim=1 with Wishart prior
## iid2d Gaussian random effect in dim=2 with Wishart prior
## iid3d Gaussian random effect in dim=3 with Wishart prior
## iid4d Gaussian random effect in dim=4 with Wishart prior
## iid5d Gaussian random effect in dim=5 with Wishart prior
## iidkd Gaussian random effect in dim=k with Wishart prior
## intslope Intecept-slope model with Wishart-prior
## linear Alternative interface to an fixed effect
## log1exp A nonlinear model of a covariate
## logdist A nonlinear model of a covariate
## matern2d Matern covariance function on a regular grid
## meb Berkson measurement error model
## mec Classical measurement error model
## ou The Ornstein-Uhlenbeck process
## prw2 Proper random walk of order 2
## revsigm Reverse sigmoidal effect of a covariate
## rgeneric Generic latent model specified using R
## rw1 Random walk of order 1
## rw2 Random walk of order 2
## rw2d Thin-plate spline model
## rw2diid Thin-plate spline with iid noise
## scopy Create a scopy of a model component
## seasonal Seasonal model for time series
## sigm Sigmoidal effect of a covariate
## slm Spatial lag model
## spde A SPDE model
## spde2 A SPDE2 model
## spde3 A SPDE3 model
## z The z-model in a classical mixed model formulation
## Section [likelihood]
## 0binomial New 0-inflated Binomial
## 0binomialS New 0-inflated Binomial Swap
## 0poisson New 0-inflated Poisson
## 0poissonS New 0-inflated Poisson Swap
## 1poisson New 1-inflated Poisson
## 1poissonS New 1-inflated Poisson Swap
## agaussian The aggregated Gaussian likelihoood
## bcgaussian The Box-Cox Gaussian likelihoood
## bell The Bell likelihood
## beta The Beta likelihood
## betabinomial The Beta-Binomial likelihood
## betabinomialna The Beta-Binomial Normal approximation likelihood
## bgev The blended Generalized Extreme Value likelihood
## binomial The Binomial likelihood
## binomialmix Binomial mixture
## cbinomial The clustered Binomial likelihood
## cennbinomial2 The CenNegBinomial2 likelihood (similar to cenpoisson2)
## cenpoisson Then censored Poisson likelihood
## cenpoisson2 Then censored Poisson likelihood (version 2)
## circularnormal The circular Gaussian likelihoood
## cloglike User-defined likelihood
## coxph Cox-proportional hazard likelihood
## dgompertzsurv destructive gompertz (survival) distribution
## dgp Discrete generalized Pareto likelihood
## egp Exteneded Generalized Pareto likelihood
## exponential The Exponential likelihood
## exponentialsurv The Exponential likelihood (survival)
## exppower The exponential power likelihoood
## fl The fl likelihood
## fmri fmri distribution (special nc-chi)
## fmrisurv fmri distribution (special nc-chi)
## gamma The Gamma likelihood
## gammacount A Gamma generalisation of the Poisson likelihood
## gammajw A special case of the Gamma likelihood
## gammajwsurv A special case of the Gamma likelihood (survival)
## gammasurv The Gamma likelihood (survival)
## gammasv The Gamma likelihood with constant rate
## gaussian The Gaussian likelihoood
## gaussianjw The GaussianJW likelihoood
## gev The Generalized Extreme Value likelihood
## ggaussian Generalized Gaussian
## ggaussianS Generalized GaussianS
## gompertz gompertz distribution
## gompertzsurv gompertz distribution
## gp Generalized Pareto likelihood
## gpoisson The generalized Poisson likelihood
## iidgamma (experimental)
## iidlogitbeta (experimental)
## lavm Link adjusted von Mises circular distribution
## loggammafrailty (experimental)
## logistic The Logistic likelihoood
## loglogistic The loglogistic likelihood
## loglogisticsurv The loglogistic likelihood (survival)
## lognormal The log-Normal likelihood
## lognormalsurv The log-Normal likelihood (survival)
## logperiodogram Likelihood for the log-periodogram
## mgamma The modal Gamma likelihood
## mgammasurv The modal Gamma likelihood (survival)
## nbinomial The negBinomial likelihood
## nbinomial2 The negBinomial2 likelihood
## nmix Binomial-Poisson mixture
## nmixnb NegBinomial-Poisson mixture
## npoisson The Normal approximation to the Poisson likelihood
## nvm Normal approx of the von Mises circular distribution
## nzpoisson The nzPoisson likelihood
## obeta The ordered Beta likelihood
## occupancy Occupancy likelihood
## poisson The Poisson likelihood
## poisson.special1 The Poisson.special1 likelihood
## pom Likelihood for the proportional odds model
## qkumar A quantile version of the Kumar likelihood
## qloglogistic A quantile loglogistic likelihood
## qloglogisticsurv A quantile loglogistic likelihood (survival)
## rcpoisson Randomly censored Poisson
## sem The SEM likelihoood
## simplex The simplex likelihood
## sn The Skew-Normal likelihoood
## stdgaussian The stdGaussian likelihoood
## stochvol The Gaussian stochvol likelihood
## stochvolln The Log-Normal stochvol likelihood
## stochvolnig The Normal inverse Gaussian stochvol likelihood
## stochvolsn The SkewNormal stochvol likelihood
## stochvolt The Student-t stochvol likelihood
## t Student-t likelihood
## tpoisson Thinned Poisson
## tstrata A stratified version of the Student-t likelihood
## tweedie Tweedie distribution
## vm von Mises circular distribution
## weibull The Weibull likelihood
## weibullsurv The Weibull likelihood (survival)
## wrappedcauchy The wrapped Cauchy likelihoood
## xbinomial The Binomial likelihood (experimental version)
## xpoisson The Poisson likelihood (expert version)
## zeroinflatedbetabinomial0 Zero-inflated Beta-Binomial, type 0
## zeroinflatedbetabinomial1 Zero-inflated Beta-Binomial, type 1
## zeroinflatedbetabinomial2 Zero inflated Beta-Binomial, type 2
## zeroinflatedbinomial0 Zero-inflated Binomial, type 0
## zeroinflatedbinomial1 Zero-inflated Binomial, type 1
## zeroinflatedbinomial2 Zero-inflated Binomial, type 2
## zeroinflatedcenpoisson0 Zero-inflated censored Poisson, type 0
## zeroinflatedcenpoisson1 Zero-inflated censored Poisson, type 1
## zeroinflatednbinomial0 Zero inflated negBinomial, type 0
## zeroinflatednbinomial1 Zero inflated negBinomial, type 1
## zeroinflatednbinomial1strata2 Zero inflated negBinomial, type 1, strata 2
## zeroinflatednbinomial1strata3 Zero inflated negBinomial, type 1, strata 3
## zeroinflatednbinomial2 Zero inflated negBinomial, type 2
## zeroinflatedpoisson0 Zero-inflated Poisson, type 0
## zeroinflatedpoisson1 Zero-inflated Poisson, type 1
## zeroinflatedpoisson2 Zero-inflated Poisson, type 2
## zeroninflatedbinomial2 Zero and N inflated binomial, type 2
## zeroninflatedbinomial3 Zero and N inflated binomial, type 3
## Section [link]
## cauchit The cauchit-link
## ccloglog The complement complementary log-log link
## cgevit Complement GEVIT link
## cloglog The complementary log-log link
## default The default link
## gevit GEVIT link
## identity The identity link
## inverse The inverse link
## log The log-link
## loga The loga-link
## logit The logit-link
## logitoffset Logit-link with an offset
## loglog The log-log link
## logoffset Log-link with an offset
## neglog The negative log-link
## powerlogit Power logit link
## pquantile The population quantile-link
## probit The probit-link
## quantile The quantile-link
## robit Robit link
## sn Skew-normal link
## special1 A special1-link function (experimental)
## special2 A special2-link function (experimental)
## sslogit Logit link with sensitivity and specificity
## tan The tan-link
## tanpi The tanpi-link
## test1 A test1-link function (experimental)
## Section [lp.scale]
## NA Section [mix]
## gaussian Gaussian mixture
## loggamma LogGamma mixture
## mloggamma Minus-LogGamma mixture
## Section [predictor]
## predictor (do not use)
## Section [prior]
## betacorrelation Beta prior for the correlation
## dirichlet Dirichlet prior
## expression: A generic prior defined using expressions
## flat A constant prior
## gamma Gamma prior
## gaussian Gaussian prior
## invalid Void prior
## jeffreystdf Jeffreys prior for the doc
## laplace Laplace prior
## linksnintercept Skew-normal-link intercept-prior
## logflat A constant prior for log(theta)
## loggamma Log-Gamma prior
## logiflat A constant prior for log(1/theta)
## logitbeta Logit prior for a probability
## logtgaussian Truncated Gaussian prior
## logtnormal Truncated Normal prior
## minuslogsqrtruncnormal (obsolete)
## mvnorm A multivariate Normal prior
## none No prior
## normal Normal prior
## pc Generic PC prior
## pc.alphaw PC prior for alpha in Weibull
## pc.ar PC prior for the AR(p) model
## pc.cor0 PC prior correlation, basemodel cor=0
## pc.cor1 PC prior correlation, basemodel cor=1
## pc.dof PC prior for log(dof-2)
## pc.egptail PC prior for the tail in the EGP likelihood
## pc.fgnh PC prior for the Hurst parameter in FGN
## pc.gamma PC prior for a Gamma parameter
## pc.gammacount PC prior for the GammaCount likelihood
## pc.gevtail PC prior for the tail in the GEV likelihood
## pc.matern PC prior for the Matern SPDE
## pc.mgamma PC prior for a Gamma parameter
## pc.prec PC prior for log(precision)
## pc.prw2.range PCprior for the range in PRW2
## pc.range PC prior for the range in the Matern SPDE
## pc.sn PC prior for the skew-normal
## pc.spde.GA (experimental)
## pom #classes-dependent prior for the POM model
## ref.ar Reference prior for the AR(p) model, p<=3
## rprior: A R-function defining the prior
## table: A generic tabulated prior
## wishart1d Wishart prior dim=1
## wishart2d Wishart prior dim=2
## wishart3d Wishart prior dim=3
## wishart4d Wishart prior dim=4
## wishart5d Wishart prior dim=5
## wishartkd Wishart prior
## Section [scopy]
## rw1 Random walk of order 1
## rw2 Random walk of order 2
## Section [wrapper]
## joint (experimental)Copying a model
We often encounter the situation where an element of a model is needed more than once for each observation. One example is where
y = a + b*w + ... for fixed weights and where is bivariate Normal and all 2-vectors are independent.
Using the model
f(idx, model="iid2d", n=2*m, ...)provide a random vector , say, with length stored as To implement this, we simply create an indentical copy of , , where (nearly). Using the copy-feature, we can do
idx = 1:m
idx.copy = m + 1:m
formula = y ~ f(idx, model="iid2d", n=2*m) + f(idx.copy, w, copy="idx") + ....recalling that the first elements is and the last elements are , and where are the weights.
The second terms define itself as a copy of , and it inherit some of its features, like and .
A copied model may also have an unknown scaling (hyperparameter), which is default fixed to be 1. In the following example, we will use this feature to estimate the unknown scaling (in this case, scaling is 2), assuming we know the precision for .
n=1000
i=1:n
j = i
z = rnorm(n)
w = runif(n)
y = z + 2*z*w + rnorm(n)
formula = y ~ f(i, model="iid",initial=0, fixed=T) +
f(j, w, copy="i", fixed=FALSE)
r = inla(formula, data = data.frame(i,j,w,y))
summary(r)## Time used:
## Pre = 0.368, Running = 1.18, Post = 0.0308, Total = 1.58
## Fixed effects:
## mean sd 0.025quant 0.5quant 0.975quant mode kld
## (Intercept) -0.175 0.059 -0.292 -0.175 -0.059 -0.175 0
##
## Random effects:
## Name Model
## i IID model
## j Copy
##
## Model hyperparameters:
## mean sd 0.025quant 0.5quant
## Precision for the Gaussian observations 23564.14 2.57e+04 1833.45 15817.16
## Beta for j 2.31 9.30e-02 2.13 2.31
## 0.975quant mode
## Precision for the Gaussian observations 91108.03 5043.58
## Beta for j 2.49 2.30
##
## Marginal log-Likelihood: -2206.90
## is computed
## Posterior summaries for the linear predictor and the fitted values are computed
## (Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')If the scaling parameter is within given range, then option \verb|range = c(low, high)|, can be given. In this case
and the prior is defined on .
If or , then the identity mapping is used. If and |verb|low!=Inf|, then the mapping is used. The case low=Inf and high!=Inf is not yet implemented.
A model or a copied model can be copied several times. The degree of closeness of and is specified by the argument , as the precision of the noise added to to get .
Replicate a model
Independent replications of a model with the same hyperparmeters can be defined using the argument ,
f(idx, model = .., replicate = r)Here, is a vector of the same length as . In this case, we use a two-dimensional index to index this (sub-)model: , so identify the first value of the second replication of this model (component). Number of replications are defined as , unless it is defined by the argument .
One example is the model `iid’:
i = 1:n
formula = y ~ f(i, model = "iid") + ...which has an alternative equivalent formulation as `n’ replications of an iid-model with length 1
In the following example, we estimate the parameters in three AR(1) time-series with the same hyperparameters (ie its replicated) but with separate means:
n = 100
y1 = arima.sim(n=n, model=list(ar=c(0.9)))+10
y2 = arima.sim(n=n, model=list(ar=c(0.9)))+20
y3 = arima.sim(n=n, model=list(ar=c(0.9)))+30
formula = y ~ mean -1 + f(i, model="ar1", replicate=r)
y = c(y1,y2,y3)
i = rep(1:n, 3)
r = rep(1:3, each=n)
mean = as.factor(r)
result = inla(formula, family = "gaussian",
data = data.frame(y, i, mean),
control.family = list(initial = 12, fixed=TRUE))
summary(result)## Time used:
## Pre = 0.135, Running = 0.298, Post = 0.0158, Total = 0.449
## Fixed effects:
## mean sd 0.025quant 0.5quant 0.975quant mode kld
## mean1 10.031 0.793 8.490 10.016 11.663 10.019 0
## mean2 20.450 0.790 18.872 20.450 22.033 20.450 0
## mean3 28.678 0.794 27.039 28.695 30.215 28.692 0
##
## Random effects:
## Name Model
## i AR1 model
##
## Model hyperparameters:
## mean sd 0.025quant 0.5quant 0.975quant mode
## Precision for i 0.255 0.057 0.153 0.252 0.375 0.250
## Rho for i 0.875 0.028 0.818 0.877 0.926 0.877
##
## Marginal log-Likelihood: -441.53
## is computed
## Posterior summaries for the linear predictor and the fitted values are computed
## (Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')All other arguments is interpreted for the basic model and also replicated. Like argument , is interpreted as each replication sums to zero, and additional constraints are also replicated.
Models with more than one type of likelihood
There is no constraint in INLA that the type of likelihood must be the same for all observations. In fact, every observation could have its own likelihood. Extentions include more than one familily, like the Normal, Poisson, etc, but also having in the model groups of observations with separate hyperparameters within each group, where the family, for example, can be the same.
In the formula
y ~ a + 1we allow to be a matrix. In this case each column of define one likelihood where the family is the same the hyperparameters are the same. For each row, only one of the columns could (but don’t have to) have an observation (non- value), the other colums must have value . All other parameters to the likelihood, like , and are used as appropriate. Example: If row column is a Poission observation, then is used as the scaling. Similar with the others. This works as only one column for each row is non-.
The argument is in the case where is a matrix, a list of families. The argument is then a list of lists; one for each family.
The first example, is a simple linear regression, where the first half of the data is observed with unknown precision (with a ‘default’ noninformative prior) and the second half of the data is observed with unknown precision . Otherwise, the two models have the same form for the linear predictor.
## Simple linear regression with observations with two different
## variances.
n = 100
N = 2*n
y = numeric(N)
x = rnorm(N)
y[1:n] = 1 + x[1:n] + rnorm(n, sd = 1/sqrt(1))
y[1:n + n] = 1 + x[1:n + n] + rnorm(n, sd = 1/sqrt(2))
Y = matrix(NA, N, 2)
Y[1:n, 1] = y[1:n]
Y[1:n + n, 2] = y[1:n + n]
formula = Y ~ x + 1
result = inla(
formula,
data = list(Y=Y, x=x),
family = c("gaussian", "gaussian"),
control.family = list(list(prior = "flat", param = numeric()),
list()))
summary(result)## Time used:
## Pre = 0.124, Running = 0.192, Post = 0.0113, Total = 0.327
## Fixed effects:
## mean sd 0.025quant 0.5quant 0.975quant mode kld
## (Intercept) 1.071 0.056 0.960 1.071 1.182 1.071 0
## x 1.073 0.057 0.962 1.073 1.185 1.073 0
##
## Model hyperparameters:
## mean sd 0.025quant 0.5quant
## Precision for the Gaussian observations 1.01 0.143 0.75 0.999
## Precision for the Gaussian observations[2] 2.20 0.311 1.64 2.180
## 0.975quant mode
## Precision for the Gaussian observations 1.31 0.985
## Precision for the Gaussian observations[2] 2.86 2.152
##
## Marginal log-Likelihood: -263.61
## is computed
## Posterior summaries for the linear predictor and the fitted values are computed
## (Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')The second example shows how to use information from two sources to estimate the effect of the covariate .
## Simple example with two types of likelihoods
n = 10
N = 2*n
## common covariates
x = rnorm(n)
## Poisson, depends on x
E1 = runif(n)
y1 = rpois(n, lambda = E1*exp(x))
## Binomial, depends on x
size = sample(1:10, size=n, replace=TRUE)
prob = exp(x)/(1+exp(x))
y2 = rbinom(n, size= size, prob = prob)
## Join them together
Y = matrix(NA, N, 2)
Y[1:n, 1] = y1
Y[1:n + n, 2] = y2
## The E for the Poisson
E = numeric(N)
E[1:n] = E1
E[1:n + n] = NA
## Ntrials for the Binomial
Ntrials = numeric(N)
Ntrials[1:n] = NA
Ntrials[1:n + n] = size
## Duplicate the covariate which is shared
X = numeric(N)
X[1:n] = x
X[1:n + n] = x
## Formula involving Y as a matrix
formula = Y ~ X - 1
## `family' is now
result = inla(formula,
family = c("poisson", "binomial"),
control.family=list(list(), list()),
data = list(Y=Y, X=X),
E = E, Ntrials = Ntrials)
summary(result)## Time used:
## Pre = 0.118, Running = 0.159, Post = 0.0059, Total = 0.283
## Fixed effects:
## mean sd 0.025quant 0.5quant 0.975quant mode kld
## X 1.139 0.292 0.567 1.138 1.713 NA 0
##
## Marginal log-Likelihood: -25.93
## is computed
## Posterior summaries for the linear predictor and the fitted values are computed
## (Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')If the covariate ‘x’ is different for the two families, and , say, then we only need to make the following changes
X = numeric(N)
X[1:n] = x
X[1:n + n] = NA
XX = numeric(N)
XX[1:n] = NA
XX[1:n + n] = xx
formula = Y ~ X + XX -1and add into the data.frame. Note how we can express the joint model as a ‘union’ of models with the use of ’s to remove terms.
In the next example, we use also the feature to estimate three replicated AR(1) models with the same hyperparamters, each observed differently.
## An example with three independent AR(1)'s with separate means, but
## with the same hyperparameters. These are observed with three
## different likelihoods.
n = 100
x1 = arima.sim(n=n, model=list(ar=c(0.9))) + 0
x2 = arima.sim(n=n, model=list(ar=c(0.9))) + 1
x3 = arima.sim(n=n, model=list(ar=c(0.9))) + 2
## Binomial observations
Nt = 10 + rpois(n,lambda=1)
y1 = rbinom(n, size=Nt, prob = exp(x1)/(1+exp(x1)))
## Poisson observations
Ep = runif(n, min=1, max=10)
y2 = rpois(n, lambda = Ep*exp(x2))
## Gaussian observations
y3 = rnorm(n, mean=x3, sd=0.1)
## stack these in a 3-column matrix with NA's where not observed
y = matrix(NA, 3*n, 3)
y[1:n, 1] = y1
y[n + 1:n, 2] = y2
y[2*n + 1:n, 3] = y3
## define the model
r = c(rep(1,n), rep(2,n), rep(3,n))
rf = as.factor(r)
i = rep(1:n, 3)
formula = y ~ f(i, model="ar1", replicate=r, constr=TRUE) + rf -1
data = list(y=y, i=i, r=r, rf=rf)
## parameters for the binomial and the poisson
Ntrials = rep(NA, 3*n)
Ntrials[1:n] = Nt
E = rep(NA, 3*n)
E[1:n + n] = Ep
result = inla(formula, family = c("binomial", "poisson", "normal"),
data = data, Ntrials = Ntrials, E = E,
control.family = list(
list(),
list(),
list()))
summary(result)## Time used:
## Pre = 0.146, Running = 0.474, Post = 0.0168, Total = 0.638
## Fixed effects:
## mean sd 0.025quant 0.5quant 0.975quant mode kld
## rf1 -0.332 0.077 -0.484 -0.332 -0.182 -0.332 0
## rf2 0.469 0.051 0.366 0.470 0.566 0.470 0
## rf3 3.241 0.046 3.146 3.241 3.335 3.241 0
##
## Random effects:
## Name Model
## i AR1 model
##
## Model hyperparameters:
## mean sd 0.025quant 0.5quant
## Precision for the Gaussian observations[3] 14.184 19.095 2.656 8.571
## Precision for i 0.432 0.077 0.291 0.428
## Rho for i 0.793 0.038 0.709 0.796
## 0.975quant mode
## Precision for the Gaussian observations[3] 61.740 4.345
## Precision for i 0.594 0.425
## Rho for i 0.860 0.802
##
## Marginal log-Likelihood: -768.81
## is computed
## Posterior summaries for the linear predictor and the fitted values are computed
## (Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')Models where the response/data depends on linear combinations of the “linear predictor” (or the sum of “fixed” and “random” effects)
In some cases, the data/response might depend on a linear combination of the “linear predictor” defined in the formula, like
y ~ 1 + zthen this implies that depends on . Suppose if depends on ? Although it is possible to express this, using the tools we already have, it is more convenient to add another layer into the model. Let be a matrix, which defines new linear predictors, from , like
Here, is the ordinary linear predictor defined using the formula, and the data depends on the linear predictor . So we might express this as
meaning in short, that
This is specified by adding the -matrix, using
control.predictor=list(A=A)The argument , which might be defined in the formula as or as an argument , does have different interpretation in the case where the -matrix is used. The rule is that in the formula, goes into , whereas an argument goes into . If we write out the expressions above adding offsets, and , we get
In the case where there is no -matrix, then \verb|offset.total = offset.arg + offset.formula|.
The following example should provide more insight. You may change and , such that , or . Note that since the response has dimension and the covariates dimension , we need to use and not a . This example also illustrates the use of offset’s.
## 'm' is the number of observations of eta*, where eta* = A eta +
## offset.arg, and A is a fixed m x n matrix, and eta has length n. An
## offset in the formula goes into 'eta' whereas an offset in the
## argument of the inla-call, goes into eta*
n = 10
m = 100
offset.formula = 10+ 1:n
offset.arg = 1 + 1:m
## a covariate
z = runif(n)
## the linear predictor eta
eta = 1 + z + offset.formula
## the linear predictor eta* = A eta + offset.arg.
A = matrix(runif(n*m), m, n);
##A = inla.as.sparse(A) ## sparse is ok
## need 'as.vector', as 'Eta' will be a sparseMatrix if 'A' is sparse
## even if ncol(Eta) = 1
Eta = as.vector(A %*% eta) + offset.arg
s = 1e-6
Y = Eta + rnorm(m, sd=s)
## for a check, we can use several offsets. here, m1=-1 and p1=1, so
## they m1+p1 = 0.
r = inla(Y ~ 1+z + offset(offset.formula) + offset(m1) + offset(p1),
## The A-matrix defined here
control.predictor = list(A = A, compute=TRUE, precision = 1e6),
## we need to use a list() as the different lengths of Y
## and z
data = list(Y=Y, z=z,
m1 = rep(-1, n),
p1 = rep(1, n),
offset.formula = offset.formula,
offset.arg = offset.arg),
## this is the offset defined in the argument of inla
offset = offset.arg,
##
control.family = list(initial = log(1/s^2), fixed=TRUE))## Warning in inla.core(formula = formula, family = family, contrasts = contrasts, : The A-matrix in the predictor (see ?control.predictor) is specified
## but an intercept is in the formula. This will likely result
## in the intercept being applied multiple times in the model, and is likely
## not what you want. See ?inla.stack for more information.
## You can remove the intercept adding ``-1'' to the formula.
summary(r)## Time used:
## Pre = 0.122, Running = 0.154, Post = 0.00674, Total = 0.283
## Fixed effects:
## mean sd 0.025quant 0.5quant 0.975quant mode kld
## (Intercept) 1 0.001 0.998 1 1.002 1 0
## z 1 0.002 0.997 1 1.003 1 0
##
## Marginal log-Likelihood: -43617.60
## is computed
## Posterior summaries for the linear predictor and the fitted values are computed
## (Posterior marginals needs also 'control.compute=list(return.marginals.predictor=TRUE)')## [1] 3.444638e-06Here is a another example where the informal formula is
y = intercept + s[j] + 0.5*s[k] + noiseInstead of using the copy feature, we can implement this model using the A-matrix feature. What we do, is to first define a linear predictor being the intercept and , then we use the -matrix to ‘construct the model’.
n = 100
s = c(-1, 0, 1)
nS = length(s)
j = sample(1L:nS, n, replace=TRUE)
k = j
k[j == 1L] = 2
k[j == 2L] = 3
k[k == 3L] = 1
noise = rnorm(n, sd=0.0001)
y = 1 + s[j] + 0.5*s[k] + noise
## build the formula such that the linear predictor is the intercept
## (index 1) and the 's' term (index 2:(n+1)). then kind of
## 'construct' the model using the A-matrix.
formula = y ~ -1 + intercept + f(idx)
A = matrix(0, n, nS+1L)
for(i in 1L:n) {
A[i, 1L] = 1
A[i, 1L + j[i]] = 1
A[i, 1L + k[i]] = 0.5
}
data = list(intercept = c(1, rep(NA, nS)), idx = c(NA, 1L:nS))
result = inla(formula, data=data, control.predictor=list(A=A))
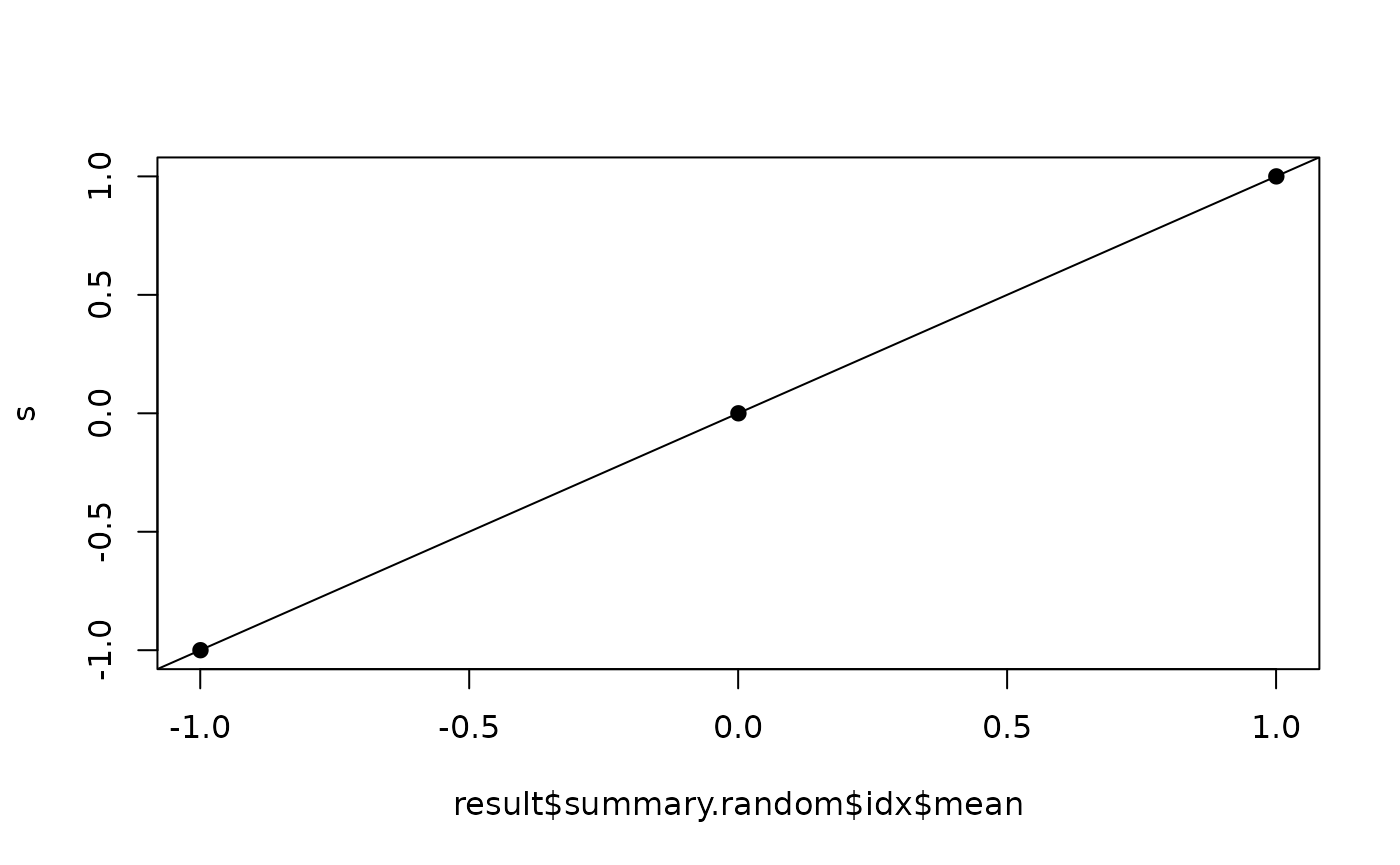
## should be a straight line
plot(result$summary.random$idx$mean, s, pch=19)
abline(a=0,b=1)