Data stacking for advanced INLA models
inla.stack.RdFunctions for combining data, effects and observation matrices into
inla.stack objects, and extracting information from such objects.
inla.stack.remove.unused(stack)
inla.stack.compress(stack, remove.unused = TRUE)
inla.stack(..., compress = TRUE, remove.unused = TRUE, multi.family = FALSE)
inla.stack.sum(
data,
A,
effects,
responses = NULL,
tag = "",
compress = TRUE,
remove.unused = TRUE
)
inla.stack.join(
...,
compress = TRUE,
remove.unused = TRUE,
multi.family = FALSE
)
inla.stack.index(stack, tag)
inla.stack.LHS(stack)
inla.stack.RHS(stack)
inla.stack.data(stack, ..., .response.name = NULL)
inla.stack.A(stack)
inla.stack.response(stack, drop = TRUE)
# S3 method for class 'inla.data.stack'
print(x, ...)Arguments
- stack
A
inla.data.stackobject, created by a call toinla.stack,inla.stack.sum, orinla.stack.join.- remove.unused
If
TRUE, compress the model by removing rows of effects corresponding to all-zero columns in theAmatrix (and removing those columns).- ...
For
inla.stack.join, two or more data stacks of classinla.data.stack, created by a call toinla.stack,inla.stack.sum, orinla.stack.join. Forinla.stack.data, a list of variables to be joined with the data list.- compress
If
TRUE, compress the model by removing duplicated rows of effects, replacing the corresponding A-matrix columns with a single column containing the sum.- multi.family
logical or character. For
inla.data.join, ifTRUE, theresponsepart of the stack is joined as alist. Ifcharacter, denotes the name of adataelement that should be joined as a multi-column matrix. Default isFALSE, which joins both thedataandresponseselements with regular row binding withdplyr::bind_rows.- data
A list or codedata.frame of named data vectors. Scalars are expanded to match the number of rows in the A matrices, or any non-scalar data vectors. An error is given if the input is inconsistent.
- A
A list of observation matrices. Scalars are expanded to diagonal matrices matching the effect vector lengths. An error is given if the input is inconsistent or ambiguous.
- effects
A collection of effects/predictors. Each list element corresponds to an observation matrix, and must either be a single vector, a list of vectors, or a
data.frame. Single-element effect vectors are expanded to vectors matching the number of columns in the corresponding A matrix. An error is given if the input is inconsistent or ombiguous.- responses
A list of response vectors, matrices, data.frame, or other special response objects, such as
inla.mdata()andinla.surv(). Each list element corresponds to one response family. In ordinary user-side code, the list has length 1, and longer lists are created by joining stacks withinla.stack(..., multi.family = TRUE).- tag
A string specifying a tag for later identification.
- .response.name
The name to assign to the response variable when extracting data from the stack. Default is
NULL, which skips the response object list.- drop
logical indicating whether to return the contained object instead of the full list, when the stack responses list has length 1. Default is
TRUE, as needed forinla()single family models. Usedrop = FALSEto extract the internal response storage, regardless of length.- x
An
inla.data.stackobject for printing
Value
A data stack of class inla.data.stack.
Elements:
data\(=(y^1, \ldots, y^p, \tilde{x}^1, \ldots, \tilde{x}^{n_k})\)A\(=\tilde{A}\)data.namesList of data names, length \(p\)effect.namesList of effect names, length \(n_k\)n.dataData length, \(n_i\)indexList indexed bytags, each element indexing into \(i=1, \ldots, n_i\)
Details
For models with a single effects collection, the outer list container for
A and effects may be omitted.
Component size definitions:
\(n_l\) effect blocks
\(n_k\) effects
\(n_i\) data values
\(n_{j,l}\) effect size for block \(l\)
\(n_j\) \(= \sum_{l=1}^{n_l} n_{j,l}\) total effect size
Input:
data\((y^1, \ldots, y^p)\) \(p\) vectors, each of length \(n_i\)
A\((A^1, \ldots, A^{n_l})\) matrices of size \(n_i \times n_{j,l}\)
effects\(\left((x^{1,1},\ldots,x^{n_k,1}), \ldots, (x^{1,n_l},\ldots,x^{n_k,n_l})\right)\) collections of effect vectors of length \(n_{j,l}\)
$$\mbox{predictor}(y^1, \ldots, y^p) \sim
\sum_{l=1}^{n_l} A^l \sum_{k=1}^{n_k} g(k, x^{k,l})
= \tilde{A} \sum_{k=1}^{n_k} g(k, \tilde{x}^k)
$$
where
$$\tilde{A} = \mbox{cbind}\left( A^1, \ldots, A^{n_l} \right)
$$
and
$$\tilde{x}^k = \mbox{rbind}\left( x^{k,1}, \ldots, x^{k,n_l} \right)
$$
and for each block \(l\), any missing
\(x^{k,l}\) is replaced by an NA vector.
Functions
inla.stack.remove.unused(): Remove unused entries from an existing stackinla.stack.compress(): Compress an existing stack by removing duplicatesinla.stack.sum(): Create data stack as a sum of predictorsinla.stack.join(): Join two or more data stacksinla.stack.index(): Extract tagged indicesinla.stack.LHS(): Extract data associated with the "left hand side" of the model (e.g. the data itself,Ntrials,link,E)inla.stack.RHS(): Extract data associated with the "right hand side" of the model (all the covariates/predictors)inla.stack.data(): Extract data for an inla call, and optionally join with other variablesinla.stack.A(): Extract the "A matrix" for control.predictorinla.stack.response(): Extract the response variable or list of response objectsprint(inla.data.stack): Print information about aninla.data.stack
Functions
inla.stack.remove.unused: Remove unused entries from an existing stackinla.stack.compress: Compress an existing stack by removing duplicatesinla.stack: Shorthand for inla.stack.join and inla.stack.suminla.stack.sum: Create data stack as a sum of predictorsinla.stack.join: Join two or more data stacksinla.stack.index: Extract tagged indicesinla.stack.LHS: Extract data associated with the "left hand side" of the model (e.g. the data itself,Ntrials,link,E)inla.stack.RHS: Extract data associated with the "right hand side" of the model (all the covariates/predictors)inla.stack.data: Extract data for an inla call, and optionally join with other variablesinla.stack.A: Extract the "A matrix" for control.predictor
See also
Examples
library(fmesher)
n <- 200
loc <- matrix(runif(n * 2), n, 2)
mesh <- fm_mesh_2d(
loc.domain = loc,
max.edge = c(0.05, 0.2)
)
proj.obs <- fm_evaluator(mesh, loc = loc)
proj.pred <- fm_evaluator(mesh, loc = mesh$loc)
spde <- inla.spde2.pcmatern(mesh,
prior.range = c(0.01, 0.01),
prior.sigma = c(10, 0.01)
)
covar <- rnorm(n)
field <- inla.qsample(n = 1, Q = inla.spde.precision(spde, theta = log(c(0.5, 1))))[, 1]
y <- 2 * covar + fm_evaluate(proj.obs, field)
A.obs <- inla.spde.make.A(mesh, loc = loc)
A.pred <- inla.spde.make.A(mesh, loc = proj.pred$loc)
stack.obs <-
inla.stack(
data = list(y = y),
A = list(A.obs, 1),
effects = list(c(
list(Intercept = 1),
inla.spde.make.index("spatial", spde$n.spde)
),
covar = covar
),
tag = "obs"
)
stack.pred <-
inla.stack(
data = list(y = NA),
A = list(A.pred),
effects = list(c(
list(Intercept = 1),
inla.spde.make.index("spatial", mesh$n)
)),
tag = "pred"
)
stack <- inla.stack(stack.obs, stack.pred)
formula <- y ~ -1 + Intercept + covar + f(spatial, model = spde)
result1 <- inla(formula,
data = inla.stack.data(stack.obs, spde = spde),
family = "gaussian",
control.predictor = list(
A = inla.stack.A(stack.obs),
compute = TRUE
)
)
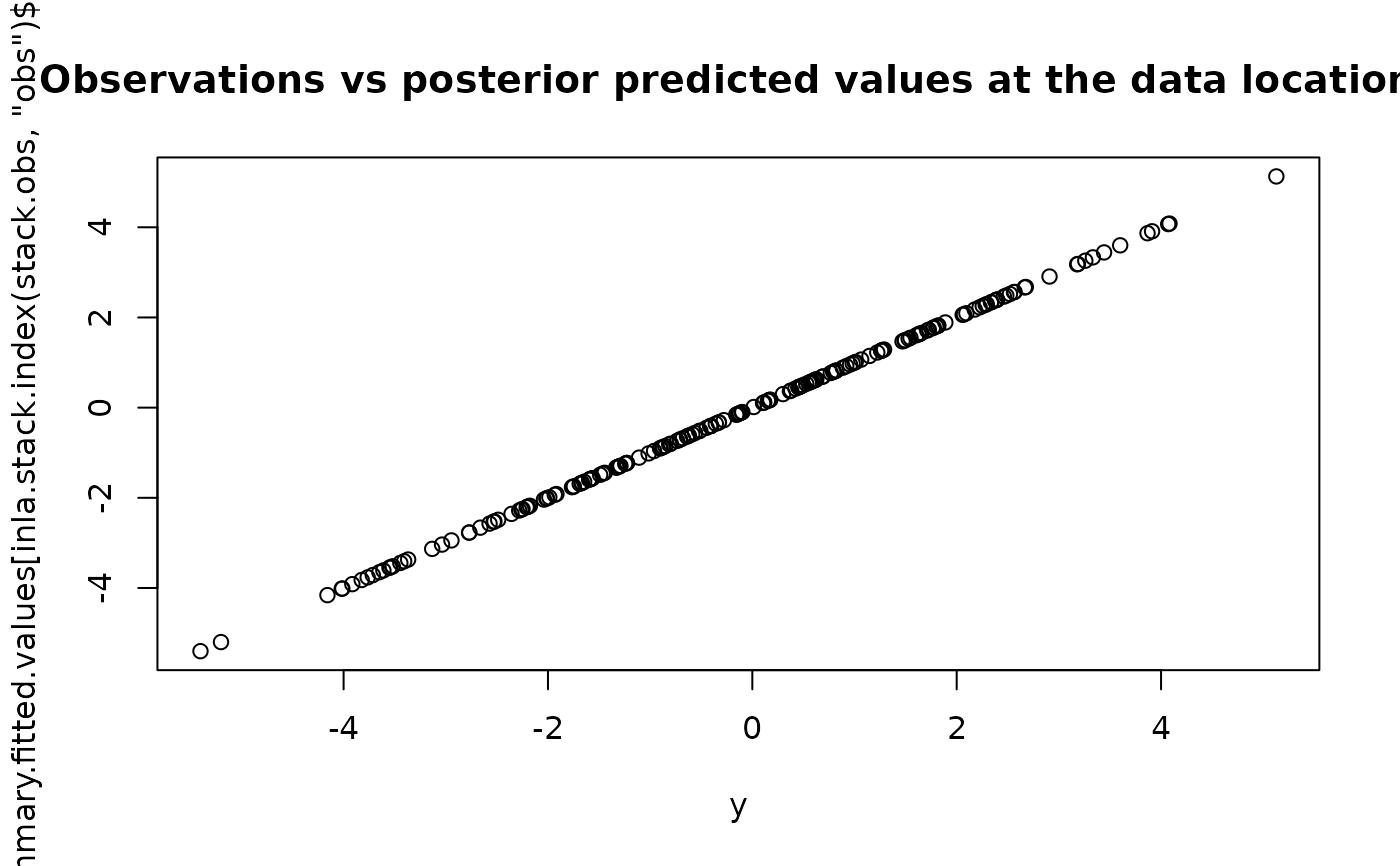
plot(y, result1$summary.fitted.values[inla.stack.index(stack.obs, "obs")$data, "mean"],
main = "Observations vs posterior predicted values at the data locations"
)
 result2 <- inla(formula,
data = inla.stack.data(stack, spde = spde),
family = "gaussian",
control.predictor = list(
A = inla.stack.A(stack),
compute = TRUE
)
)
field.pred <- fm_evaluate(
proj.pred,
result2$summary.fitted.values[inla.stack.index(stack, "pred")$data, "mean"]
)
field.pred.sd <- fm_evaluate(
proj.pred,
result2$summary.fitted.values[inla.stack.index(stack, "pred")$data, "sd"]
)
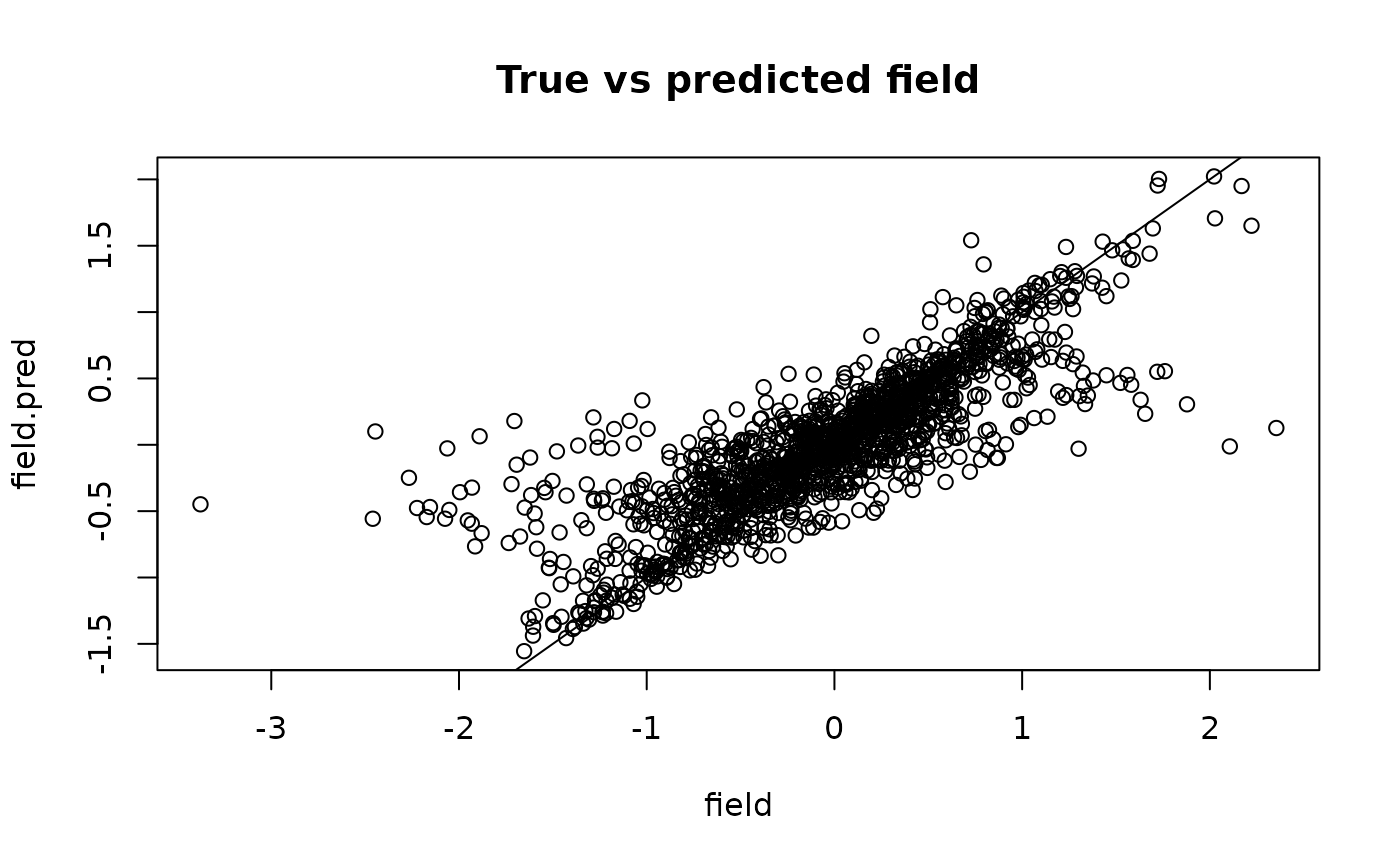
plot(field, field.pred, main = "True vs predicted field")
abline(0, 1)
result2 <- inla(formula,
data = inla.stack.data(stack, spde = spde),
family = "gaussian",
control.predictor = list(
A = inla.stack.A(stack),
compute = TRUE
)
)
field.pred <- fm_evaluate(
proj.pred,
result2$summary.fitted.values[inla.stack.index(stack, "pred")$data, "mean"]
)
field.pred.sd <- fm_evaluate(
proj.pred,
result2$summary.fitted.values[inla.stack.index(stack, "pred")$data, "sd"]
)
plot(field, field.pred, main = "True vs predicted field")
abline(0, 1)
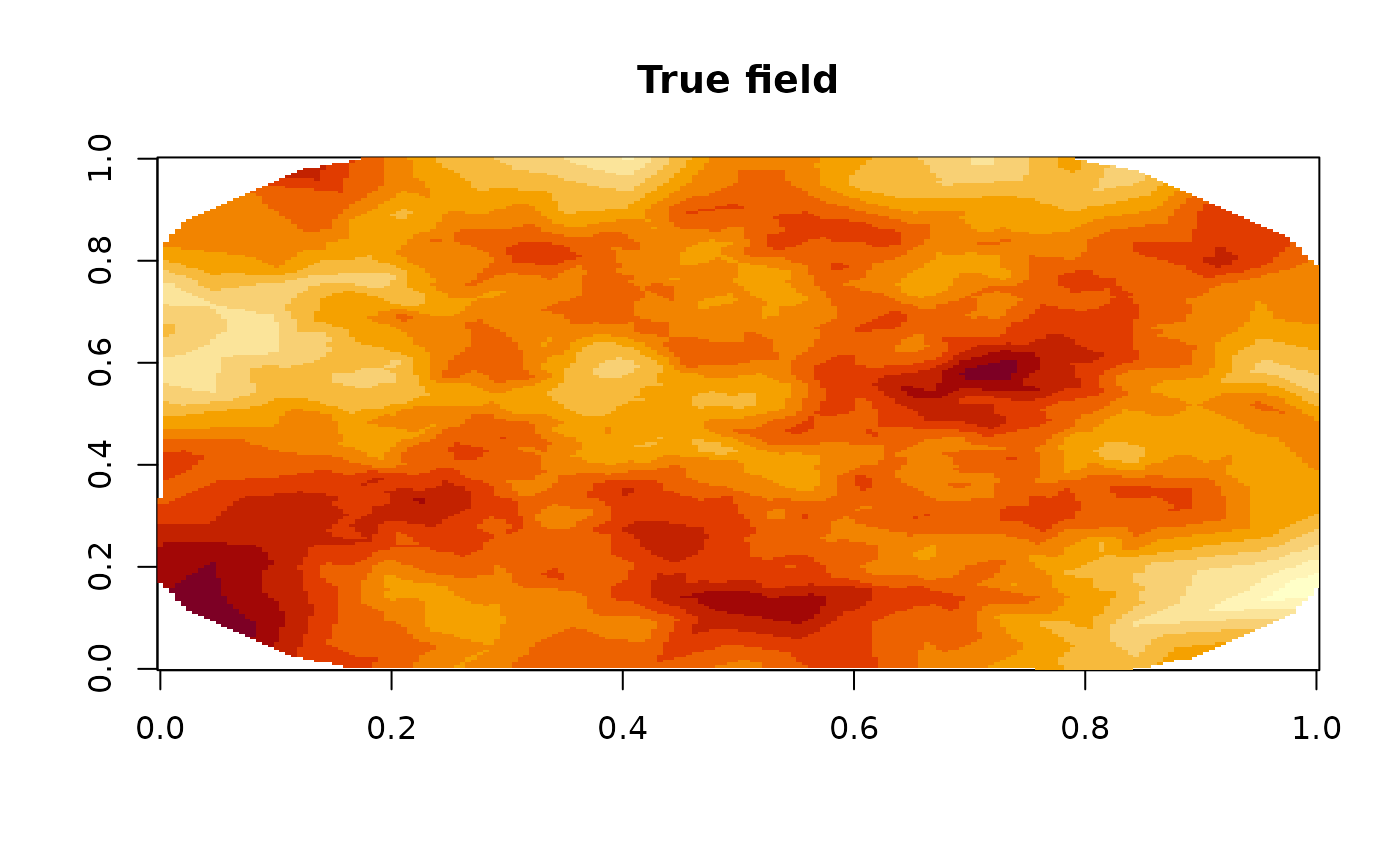
 image(fm_evaluate(mesh,
field = field,
dims = c(200, 200)
),
main = "True field"
)
image(fm_evaluate(mesh,
field = field,
dims = c(200, 200)
),
main = "True field"
)
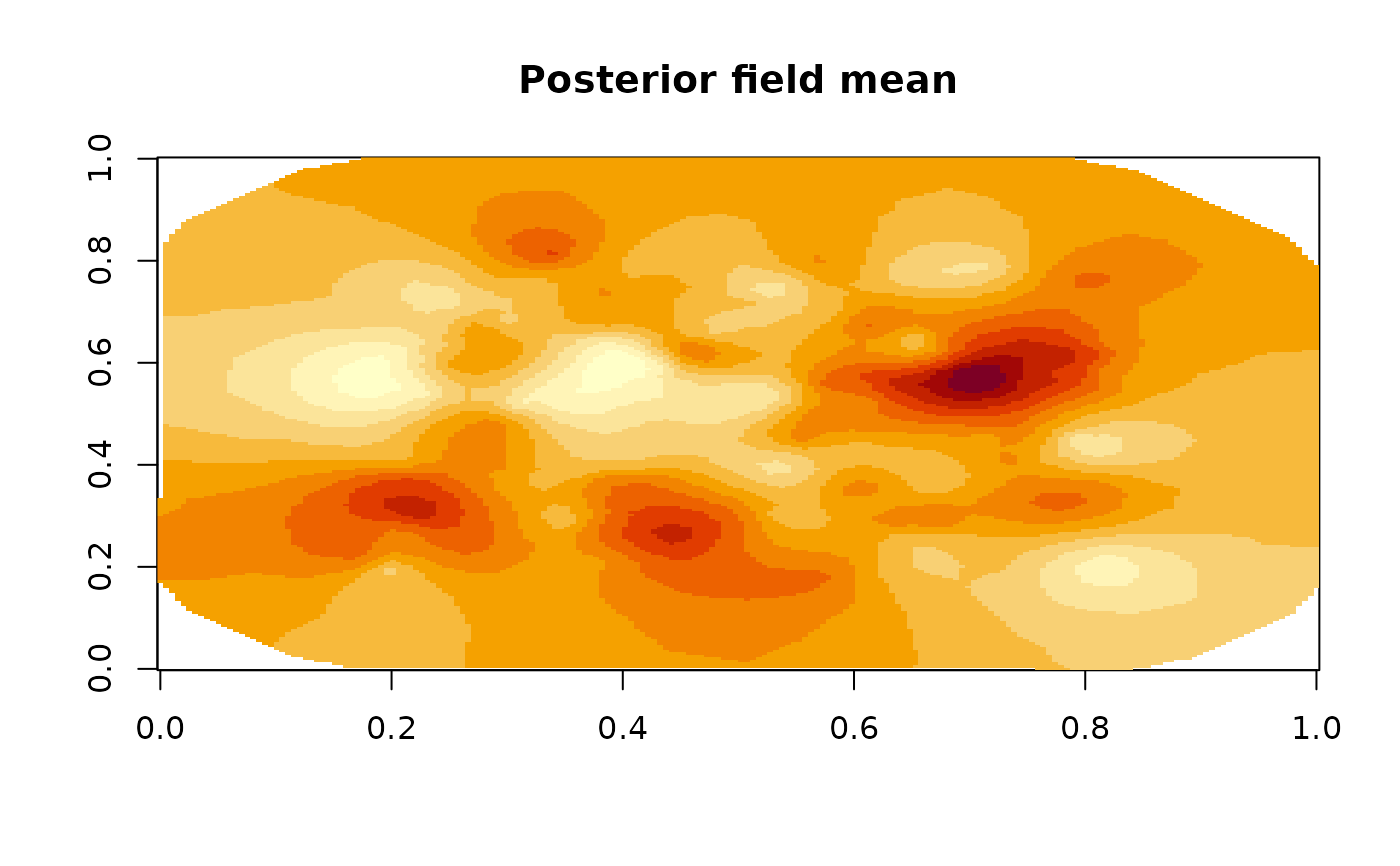
 image(fm_evaluate(mesh,
field = field.pred,
dims = c(200, 200)
),
main = "Posterior field mean"
)
image(fm_evaluate(mesh,
field = field.pred,
dims = c(200, 200)
),
main = "Posterior field mean"
)
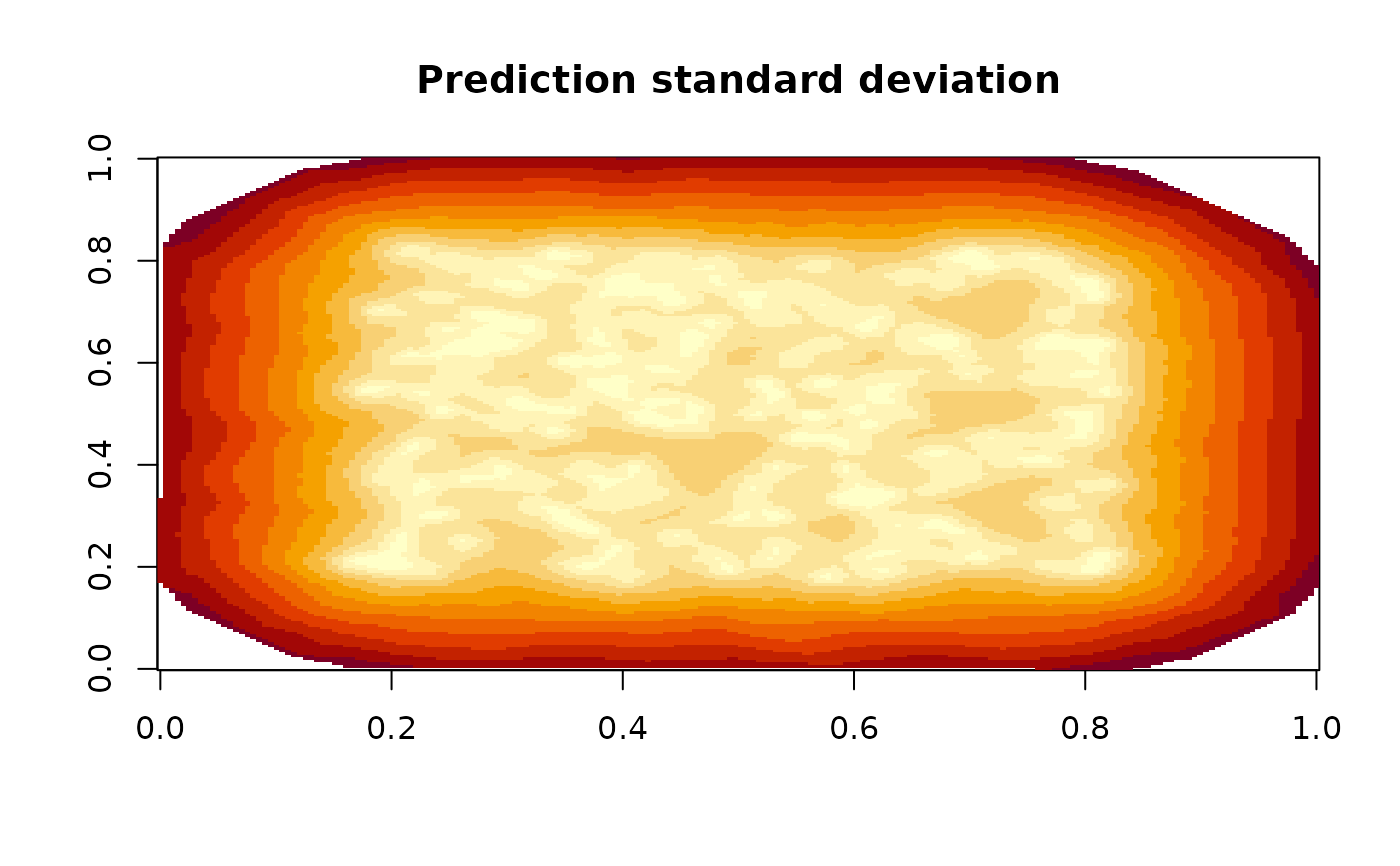
 image(fm_evaluate(mesh,
field = field.pred.sd,
dims = c(200, 200)
),
main = "Prediction standard deviation"
)
image(fm_evaluate(mesh,
field = field.pred.sd,
dims = c(200, 200)
),
main = "Prediction standard deviation"
)
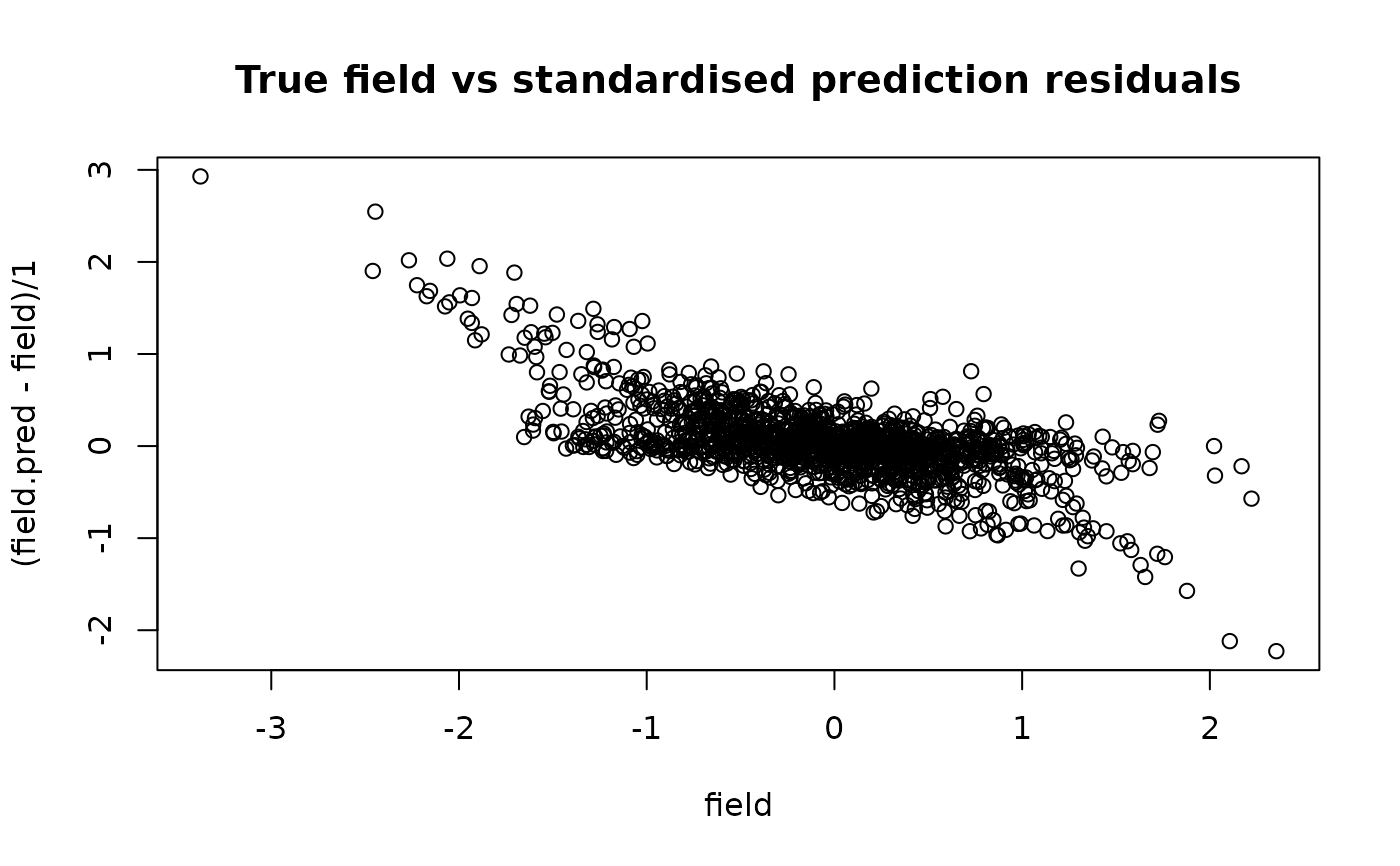
 plot(field, (field.pred - field) / 1,
main = "True field vs standardised prediction residuals"
)
plot(field, (field.pred - field) / 1,
main = "True field vs standardised prediction residuals"
)