Generate samples, and functions thereof, from an approximated posterior of a fitted model
posterior.sample.RdThis function generate samples, and functions of those, from an approximated posterior of a fitted model (an inla-object)
The hyperparameters are sampled from the configurations used to do the
numerical integration, hence if you want a higher resolution, you need to to
change the int.stratey variable and friends. The latent field is
sampled from the Gaussian approximation conditioned on the hyperparameters,
but with a correction for the mean (default), and optional (and by default)
corrected for the estimated skewness.
The log.density report is only correct when there is no constraints. With constraints, it correct the Gaussian part of the sample for the constraints.
After the sample is (optional) skewness corrected, the log.density is is not exact for correcting for constraints, but the error is very small in most cases.
inla.posterior.sample(
n = 1L,
result,
selection = list(),
intern = FALSE,
use.improved.mean = TRUE,
skew.corr = TRUE,
add.names = TRUE,
seed = 0L,
num.threads = NULL,
parallel.configs = TRUE,
verbose = FALSE
)
inla.posterior.sample.eval(fun, samples, return.matrix = TRUE, ...)Arguments
- n
Number of samples.
- result
The inla-object, ie the output from an
inla-call. Theinla-object must be created withcontrol.compute=list(config=TRUE).- selection
Select what part of the sample to return. By default, the whole sample is returned.
selectionis a named list with the name of the components of the sample, and what indices of them to return. Names includeAPredictor,Predictor,(Intercept), and otherwise names in the formula. The values of the list, is interpreted as indices. If they are negative, they are interpreted as 'not', a zero is interpreted as 'all', and positive indices are interpreted as 'only'. The names of elements of each samples refer to the indices in the full sample. NOTE THAT USING THIS ARGUMENT WILL MAKEinla.posterior.sample.evalFAIL.- intern
Logical. If
TRUEthen produce samples in the internal scale for the hyperparmater, ifFALSEthen produce samples in the user-scale. (For example log-precision (intern) and precision (user-scale))- use.improved.mean
Logical. If
TRUEthen use the marginal mean values when constructing samples. IfFALSEthen use the mean in the Gaussian approximations.- skew.corr
Logical. If
TRUEthen correct samples for skewness, ifFALSE, do not correct samples for skewness (ie use the Gaussian).- add.names
Logical. If
TRUEthen add name for each elements of each sample. IfFALSE, only add name for the first sample. (This save space.)- seed
See the same argument in
?inla.qsamplefor further information. In order to produce reproducible results, you ALSO need to make sure the RNG in R is in the same state, see example below. Whenseedis non-zero,num.threadsis forced to "1:1" and parallel.configs is set toFALSE, since parallel sampling would not produce a reproducible sequence of pseudo-random numbers.- num.threads
The number of threads to use in the format 'A:B' defining the number threads in the outer (A) and inner (B) layer for nested parallelism. A '0' will be replaced intelligently.
seed!=0requires serial comptuations.- parallel.configs
Logical. If
TRUEand not on Windows, then try to run each configuration in parallel (not Windows) usingAthreads (seenum.threads), where each of them is usingB:0threads.- verbose
Logical. Run in verbose mode or not.
- fun
The function to evaluate for each sample. Upon entry, the variable names defined in the model are defined as the value of the sample. The list of names are defined in
result$misc$configs$contentswhereresultis aninla-object. This includes predefined names for for the linear predictor (PredictorandAPredictor), and the intercept ((Intercept)orIntercept). The hyperparameters are defined astheta, no matter if they are in the internal scale or not. The functionfuncan also return a vector. To simplify usage,funcan also be a vector character's. In this casefunit is interpreted as (strict) variable names, and a function is created that return these variables: if argumentfunequalsc("Intercept", "a[1:2]"), then this is equivalent to passfunction() return(c(get('Intercept'), get('a[1:2]'))).- samples
samplesis the output frominla.posterior.sample()- return.matrix
Logical. If
TRUE, then return the samples offunas matrix, otherwise, as a list.- ...
Additional arguments to
fun
Value
inla.posterior.sample returns a list of the samples, where
each sample is a list with names hyperpar and latent, and with
their marginal densities in logdens$hyperpar and
logdens$latent and the joint density is in logdens$joint.
inla.posterior.sample.eval return a list or a matrix of fun
applied to each sample.
Examples
r = inla(y ~ 1 ,data = data.frame(y=rnorm(1)), control.compute = list(config=TRUE))
samples = inla.posterior.sample(2,r)
## reproducible results:
inla.seed = as.integer(runif(1)*.Machine$integer.max)
set.seed(12345)
x = inla.posterior.sample(10, r, seed = inla.seed, num.threads="1:1")
set.seed(12345)
xx = inla.posterior.sample(10, r, seed = inla.seed, num.threads="1.1")
all.equal(x, xx)
#> [1] TRUE
set.seed(1234)
n = 25
xx = rnorm(n)
yy = rev(xx)
z = runif(n)
y = rnorm(n)
r = inla(y ~ 1 + z + f(xx) + f(yy, copy="xx"),
data = data.frame(y, z, xx, yy),
control.compute = list(config=TRUE),
family = "gaussian")
r.samples = inla.posterior.sample(10, r)
fun = function(...) {
mean(xx) - mean(yy)
}
f1 = inla.posterior.sample.eval(fun, r.samples)
fun = function(...) {
c(exp(Intercept), exp(Intercept + z))
}
f2 = inla.posterior.sample.eval(fun, r.samples)
fun = function(...) {
return (theta[1]/(theta[1] + theta[2]))
}
f3 = inla.posterior.sample.eval(fun, r.samples)
## Predicting nz new observations, and
## comparing the estimated one with the true one
set.seed(1234)
n = 100
alpha = beta = s = 1
z = rnorm(n)
y = alpha + beta * z + rnorm(n, sd = s)
r = inla(y ~ 1 + z,
data = data.frame(y, z),
control.compute = list(config=TRUE),
family = "gaussian")
r.samples = inla.posterior.sample(10^3, r)
## just return samples of the intercept
intercepts = inla.posterior.sample.eval("Intercept", r.samples)
nz = 3
znew = rnorm(nz)
fun = function(zz = NA) {
## theta[1] is the precision
return (Intercept + z * zz +
rnorm(length(zz), sd = sqrt(1/theta[1])))
}
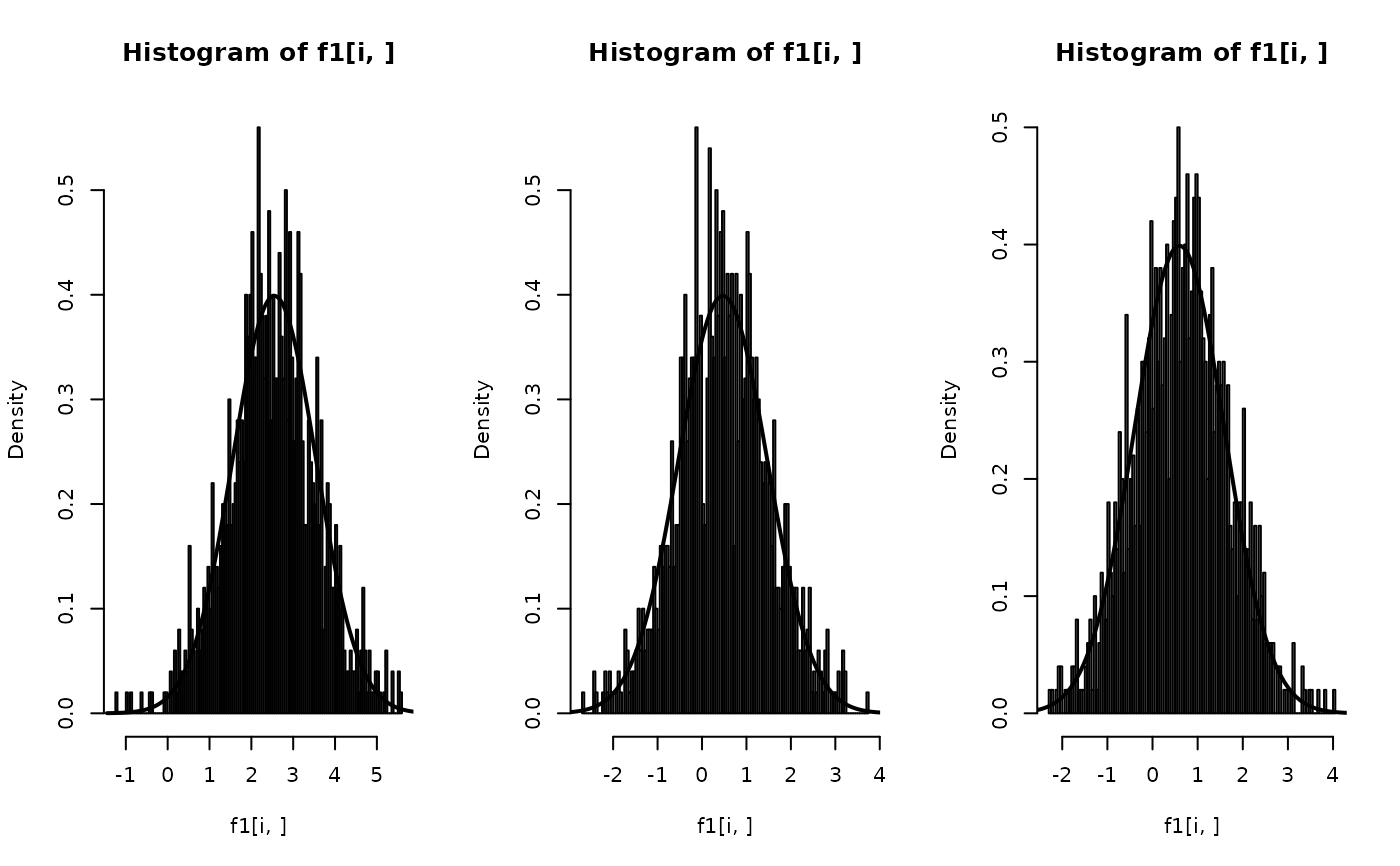
par(mfrow=c(1, nz))
f1 = inla.posterior.sample.eval(fun, r.samples, zz = znew)
for(i in 1:nz) {
hist(f1[i, ], n = 100, prob = TRUE)
m = alpha + beta * znew[i]
xx = seq(m-4*s, m+4*s, by = s/100)
lines(xx, dnorm(xx, mean=m, sd = s), lwd=2)
}
 ##
## Be aware that using non-clean variable names might be a little tricky
##
n <- 100
X <- matrix(rnorm(n^2), n, 2)
#> Warning: data length differs from size of matrix: [10000 != 100 x 2]
x <- X[, 1]
xx <- X[, 2]
xxx <- x*xx
y <- 1 + 2*x + 3*xx + 4*xxx + rnorm(n, sd = 0.01)
r <- inla(y ~ X[, 1]*X[, 2],
data = list(y = y, X = X),
control.compute = list(config = TRUE))
print(round(dig = 4, r$summary.fixed[,"mean"]))
#> [1] 1.0018 1.9995 3.0000 4.0007
sam <- inla.posterior.sample(100, r)
sam.extract <- inla.posterior.sample.eval(
(function(...) {
beta.1 <- get("X[, 1]")
beta.2 <- get("X[, 2]")
beta.12 <- get("X[, 1]:X[, 2]")
return(c(Intercept, beta.1, beta.2, beta.12))
}), sam)
print(round(dig = 4, rowMeans(sam.extract)))
#> fun[1] fun[2] fun[3] fun[4]
#> 1.0020 1.9995 3.0000 4.0006
## a simpler form can also be used here, and in the examples below
sam.extract <- inla.posterior.sample.eval(
c("Intercept", "X[, 1]", "X[, 2]", "X[, 1]:X[, 2]"), sam)
print(round(dig = 4, rowMeans(sam.extract)))
#> fun[1] fun[2] fun[3] fun[4]
#> 1.0020 1.9995 3.0000 4.0006
r <- inla(y ~ x + xx + xxx,
data = list(y = y, x = x, xx = xx, xxx = xxx),
control.compute = list(config = TRUE))
sam <- inla.posterior.sample(100, r)
sam.extract <- inla.posterior.sample.eval(
(function(...) {
return(c(Intercept, x, xx, xxx))
}), sam)
print(round(dig = 4, rowMeans(sam.extract)))
#> fun[1] fun[2] fun[3] fun[4]
#> 1.0018 1.9994 3.0000 4.0006
sam.extract <- inla.posterior.sample.eval(c("Intercept", "x", "xx", "xxx"), sam)
print(round(dig = 4, rowMeans(sam.extract)))
#> fun[1] fun[2] fun[3] fun[4]
#> 1.0018 1.9994 3.0000 4.0006
r <- inla(y ~ x*xx,
data = list(y = y, x = x, xx = xx),
control.compute = list(config = TRUE))
sam <- inla.posterior.sample(100, r)
sam.extract <- inla.posterior.sample.eval(
(function(...) {
return(c(Intercept, x, xx, get("x:xx")))
}), sam)
print(round(dig = 4, rowMeans(sam.extract)))
#> fun[1] fun[2] fun[3] fun[4]
#> 1.0017 1.9995 3.0001 4.0007
sam.extract <- inla.posterior.sample.eval(c("Intercept", "x", "xx", "x:xx"), sam)
print(round(dig = 4, rowMeans(sam.extract)))
#> fun[1] fun[2] fun[3] fun[4]
#> 1.0017 1.9995 3.0001 4.0007
##
## Be aware that using non-clean variable names might be a little tricky
##
n <- 100
X <- matrix(rnorm(n^2), n, 2)
#> Warning: data length differs from size of matrix: [10000 != 100 x 2]
x <- X[, 1]
xx <- X[, 2]
xxx <- x*xx
y <- 1 + 2*x + 3*xx + 4*xxx + rnorm(n, sd = 0.01)
r <- inla(y ~ X[, 1]*X[, 2],
data = list(y = y, X = X),
control.compute = list(config = TRUE))
print(round(dig = 4, r$summary.fixed[,"mean"]))
#> [1] 1.0018 1.9995 3.0000 4.0007
sam <- inla.posterior.sample(100, r)
sam.extract <- inla.posterior.sample.eval(
(function(...) {
beta.1 <- get("X[, 1]")
beta.2 <- get("X[, 2]")
beta.12 <- get("X[, 1]:X[, 2]")
return(c(Intercept, beta.1, beta.2, beta.12))
}), sam)
print(round(dig = 4, rowMeans(sam.extract)))
#> fun[1] fun[2] fun[3] fun[4]
#> 1.0020 1.9995 3.0000 4.0006
## a simpler form can also be used here, and in the examples below
sam.extract <- inla.posterior.sample.eval(
c("Intercept", "X[, 1]", "X[, 2]", "X[, 1]:X[, 2]"), sam)
print(round(dig = 4, rowMeans(sam.extract)))
#> fun[1] fun[2] fun[3] fun[4]
#> 1.0020 1.9995 3.0000 4.0006
r <- inla(y ~ x + xx + xxx,
data = list(y = y, x = x, xx = xx, xxx = xxx),
control.compute = list(config = TRUE))
sam <- inla.posterior.sample(100, r)
sam.extract <- inla.posterior.sample.eval(
(function(...) {
return(c(Intercept, x, xx, xxx))
}), sam)
print(round(dig = 4, rowMeans(sam.extract)))
#> fun[1] fun[2] fun[3] fun[4]
#> 1.0018 1.9994 3.0000 4.0006
sam.extract <- inla.posterior.sample.eval(c("Intercept", "x", "xx", "xxx"), sam)
print(round(dig = 4, rowMeans(sam.extract)))
#> fun[1] fun[2] fun[3] fun[4]
#> 1.0018 1.9994 3.0000 4.0006
r <- inla(y ~ x*xx,
data = list(y = y, x = x, xx = xx),
control.compute = list(config = TRUE))
sam <- inla.posterior.sample(100, r)
sam.extract <- inla.posterior.sample.eval(
(function(...) {
return(c(Intercept, x, xx, get("x:xx")))
}), sam)
print(round(dig = 4, rowMeans(sam.extract)))
#> fun[1] fun[2] fun[3] fun[4]
#> 1.0017 1.9995 3.0001 4.0007
sam.extract <- inla.posterior.sample.eval(c("Intercept", "x", "xx", "x:xx"), sam)
print(round(dig = 4, rowMeans(sam.extract)))
#> fun[1] fun[2] fun[3] fun[4]
#> 1.0017 1.9995 3.0001 4.0007