Matern SPDE model object for INLA
inla.spde2.matern.RdCreate an inla.spde2 model object for a Matern model. Use
inla.spde2.pcmatern instead for a PC prior for the parameters.
inla.spde2.matern(
mesh,
alpha = 2,
param = NULL,
constr = FALSE,
extraconstr.int = NULL,
extraconstr = NULL,
fractional.method = c("parsimonious", "null"),
B.tau = matrix(c(0, 1, 0), 1, 3),
B.kappa = matrix(c(0, 0, 1), 1, 3),
prior.variance.nominal = 1,
prior.range.nominal = NULL,
prior.tau = NULL,
prior.kappa = NULL,
theta.prior.mean = NULL,
theta.prior.prec = 0.1,
n.iid.group = 1,
...
)
inla.spde2.theta2phi0(spde, theta)
inla.spde2.theta2phi1(spde, theta)
inla.spde2.theta2phi2(spde, theta)Arguments
- mesh
The mesh to build the model on, as an
fmesher::fm_mesh_1d(),fmesher::fm_mesh_2d(),fmesher::fm_mesh_3d(), orfmesher::fm_collect()object.- alpha
Fractional operator order, \(0<\alpha\leq 2\) supported. (\(\nu=\alpha-d/2\))
- param
Parameter, e.g. generated by
param2.matern.orig- constr
If
TRUE, apply an integrate-to-zero constraint. DefaultFALSE.- extraconstr.int
Field integral constraints.
- extraconstr
Direct linear combination constraints on the basis weights.
- fractional.method
Specifies the approximation method to use for fractional (non-integer)
alphavalues.'parsimonious'gives an overall approximate minimal covariance error,'null'uses approximates low-order properties.- B.tau
Matrix with specification of log-linear model for \(\tau\).
- B.kappa
Matrix with specification of log-linear model for \(\kappa\).
- prior.variance.nominal
Nominal prior mean for the field variance
- prior.range.nominal
Nominal prior mean for the spatial range
- prior.tau
Prior mean for tau (overrides
prior.variance.nominal)- prior.kappa
Prior mean for kappa (overrides
prior.range.nominal)- theta.prior.mean
(overrides
prior.*)- theta.prior.prec
Scalar, vector or matrix, specifying the joint prior precision for \(theta\).
- n.iid.group
If greater than 1, build an explicitly iid replicated model, to support constraints applied to the combined replicates, for example in a time-replicated spatial model. Constraints can either be specified for a single mesh, in which case it's applied to the average of the replicates (
ncol(A)should bemesh$nfor 2D meshes,mesh$mfor 1D), or as general constraints on the collection of replicates (ncol(A)should bemesh$n * n.iid.groupfor 2D meshes,mesh$m * n.iid.groupfor 1D).- ...
Additional parameters for special uses.
- spde
An spde model object
- theta
Parameters in the model's internal scale
Value
An inla.spde2 object.
Details
This method constructs a Matern SPDE model, with spatial scale parameter \(\kappa(u)\) and variance rescaling parameter \(\tau(u)\).
$$(\kappa^2(u)-\Delta)^{\alpha/2}(\tau(u) $$$$ x(u))=W(u)$$
Stationary models are supported for \(0 < \alpha \leq 2\), with spectral
approximation methods used for non-integer \(\alpha\), with approximation
method determined by fractional.method.
Non-stationary models are supported for \(\alpha=2\) only, with
\(\log\tau(u) = B^\tau_0(u) + \sum_{k=1}^p B^\tau_k(u) \)\( \theta_k\)
\(\log\kappa(u) = B^{\kappa}_0(u) + \sum_{k=1}^p B^{\kappa}_k(u) \)\( \theta_k\)
The same parameterisation is used in the stationary cases, but with \(B^\tau_0\), \(B^\tau_k\), \(B^\kappa_0\), and \(B^\tau_k\) constant across \(u\).
Integration and other general linear constraints are supported via the
constr, extraconstr.int, and extraconstr parameters,
which also interact with n.iid.group.
Functions
inla.spde2.theta2phi0(): Convert from theta vector to phi0 values in the internal spde2 model representationinla.spde2.theta2phi1(): Convert from theta vector to phi1 values in the internal spde2 model representationinla.spde2.theta2phi2(): Convert from theta vector to phi2 values in the internal spde2 model representation
See also
Examples
n <- 100
field.fcn <- function(loc) (10 * cos(2 * pi * 2 * (loc[, 1] + loc[, 2])))
loc <- matrix(runif(n * 2), n, 2)
## One field, 2 observations per location
idx.y <- rep(1:n, 2)
y <- field.fcn(loc[idx.y, ]) + rnorm(length(idx.y))
mesh <- fm_rcdt_2d_inla(loc, refine = list(max.edge = 0.05))
spde <- inla.spde2.matern(mesh)
data <- list(y = y, field = mesh$idx$loc[idx.y])
formula <- y ~ -1 + f(field, model = spde)
result <- inla(formula, data = data, family = "normal")
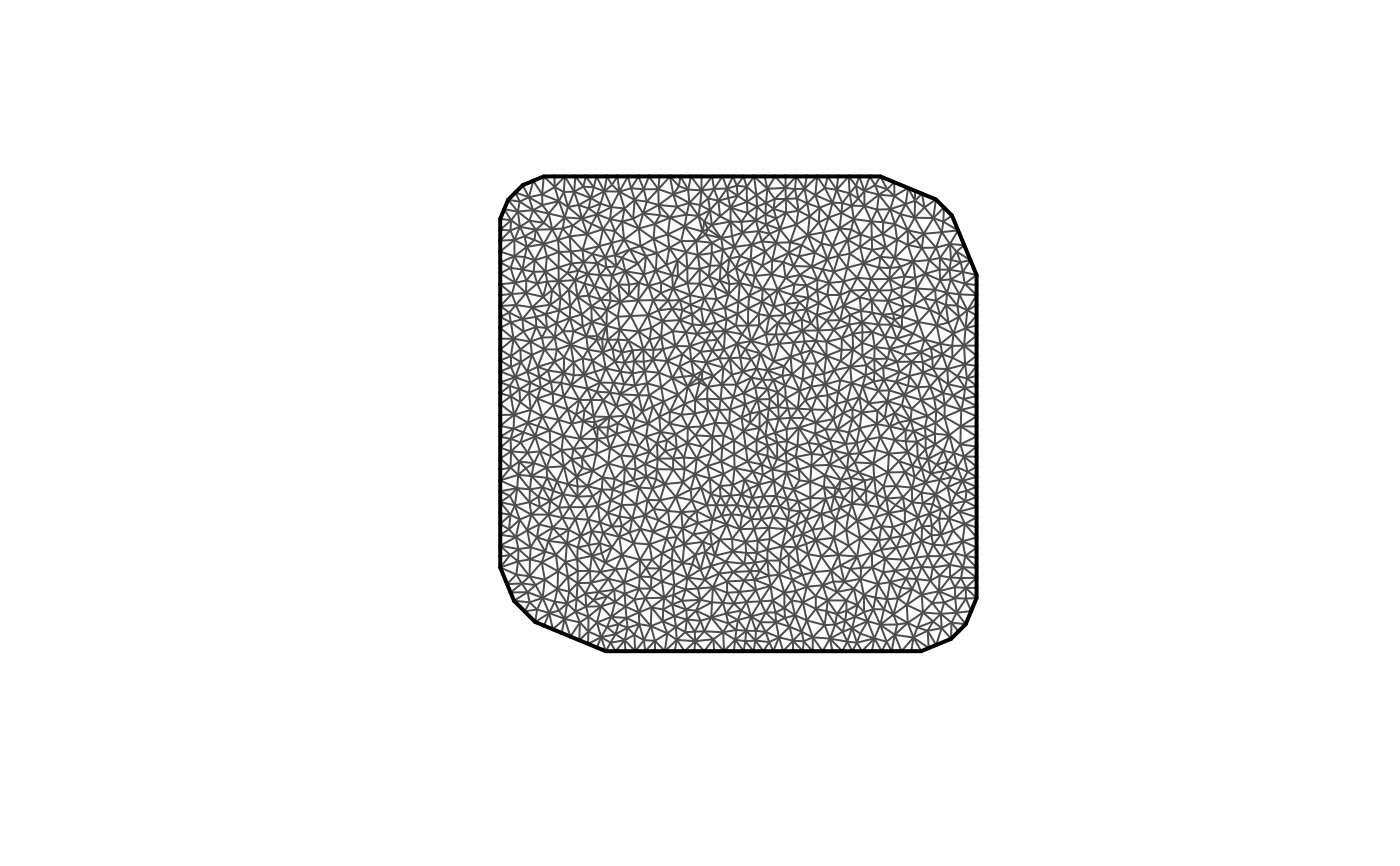
## Plot the mesh structure:
plot(mesh)
 # \donttest{
if (require(rgl)) {
col.pal <- colorRampPalette(c("blue", "cyan", "green", "yellow", "red"))
## Plot the posterior mean:
plot(mesh,
rgl = TRUE,
result$summary.random$field[, "mean"],
color.palette = col.pal
)
## Plot residual field:
plot(mesh,
rgl = TRUE,
result$summary.random$field[, "mean"] - field.fcn(mesh$loc),
color.palette = col.pal
)
}
#> Error: The `rgl` argument of `plot.fm_mesh_2d()` was deprecated in fmesher
#> 0.1.0 and is now defunct.
#> ℹ Please use `plot_rgl()` instead.
# }
result.field <- inla.spde.result(result, "field", spde)
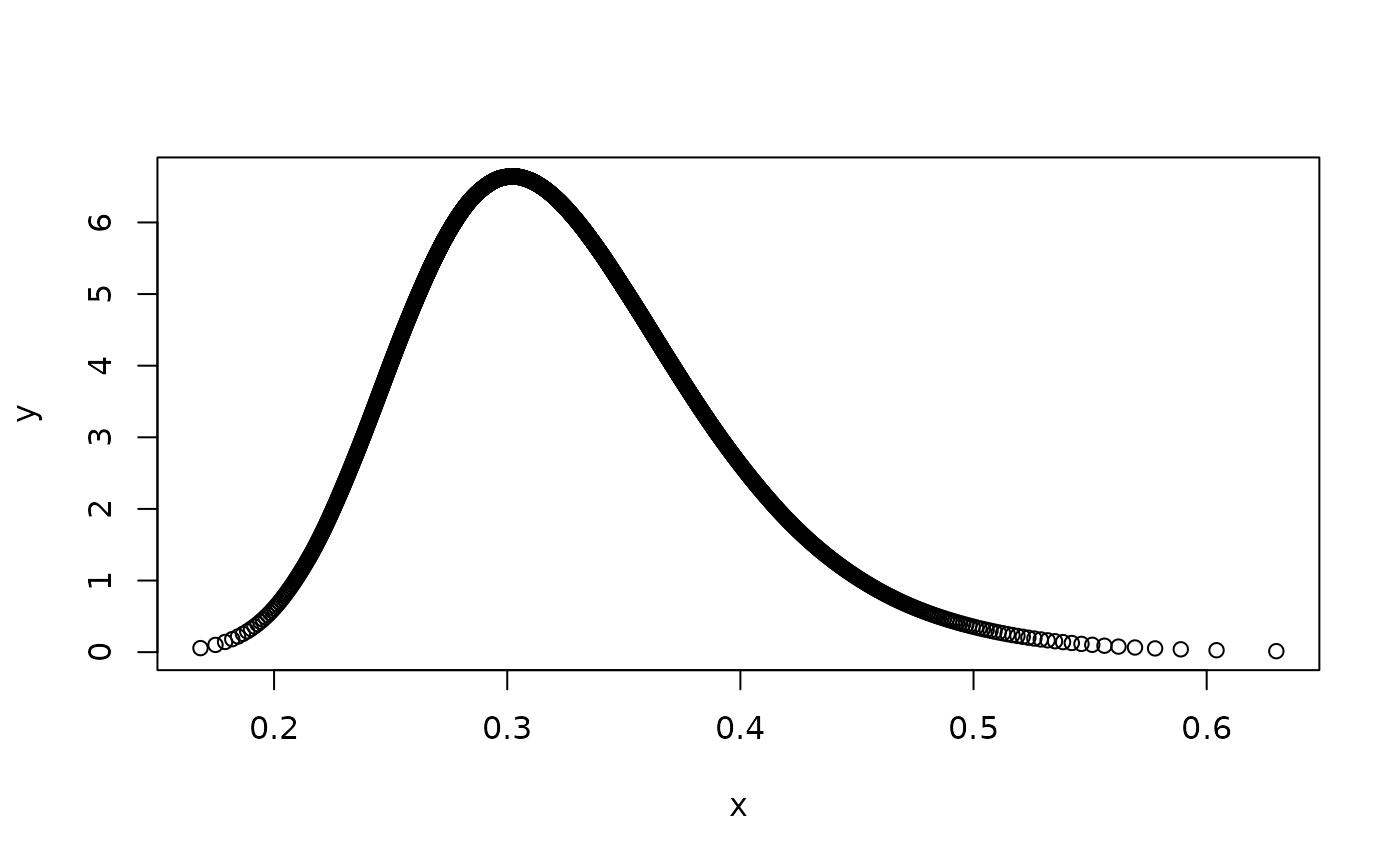
plot(result.field$marginals.range.nominal[[1]])
# \donttest{
if (require(rgl)) {
col.pal <- colorRampPalette(c("blue", "cyan", "green", "yellow", "red"))
## Plot the posterior mean:
plot(mesh,
rgl = TRUE,
result$summary.random$field[, "mean"],
color.palette = col.pal
)
## Plot residual field:
plot(mesh,
rgl = TRUE,
result$summary.random$field[, "mean"] - field.fcn(mesh$loc),
color.palette = col.pal
)
}
#> Error: The `rgl` argument of `plot.fm_mesh_2d()` was deprecated in fmesher
#> 0.1.0 and is now defunct.
#> ℹ Please use `plot_rgl()` instead.
# }
result.field <- inla.spde.result(result, "field", spde)
plot(result.field$marginals.range.nominal[[1]])